Difference between revisions of "General Information/IS91 and ISCR"
| Line 20: | Line 20: | ||
==Bibliography== | ==Bibliography== | ||
| − | < | + | {{Reflist|32em}} |
| + | <br /> | ||
| + | <hr> | ||
| + | {{TnPedia}} | ||
Revision as of 13:23, 21 March 2022
A final example of the subtle line dividing IS and transposons is found in the IS91/ISCR group (Fig.12.1). IS91 was identified some time ago[1] and carries a single Tpase orf. More recently, a group of related elements, ISCR (IS with a "Common Region") was described [reviewed in [2][3][4]] (see the IS91 and ISCR section).
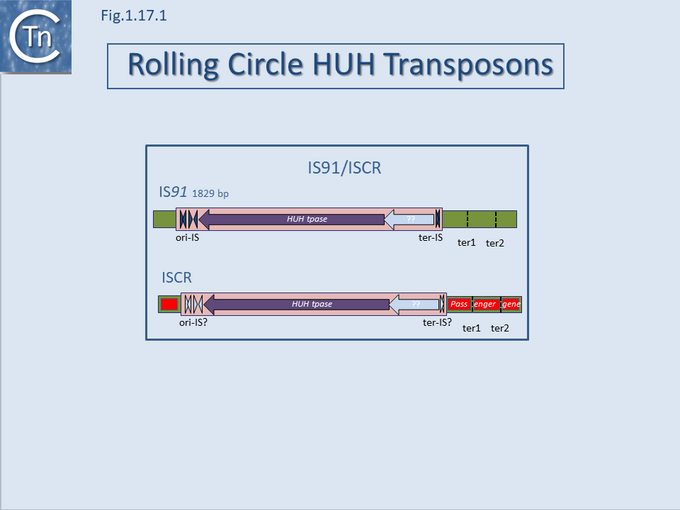
Although there has been no formal demonstration that these actually transpose, the CR is an orf which resembles the IS91 family Tpases[5]. The major feature of ISCR elements is that they are associated with a diverse variety of antibiotic resistance genes and, particularly in the case of Pseudomonas ISCR, aromatic degradation pathways, both upstream and downstream of the Tpase ORF.
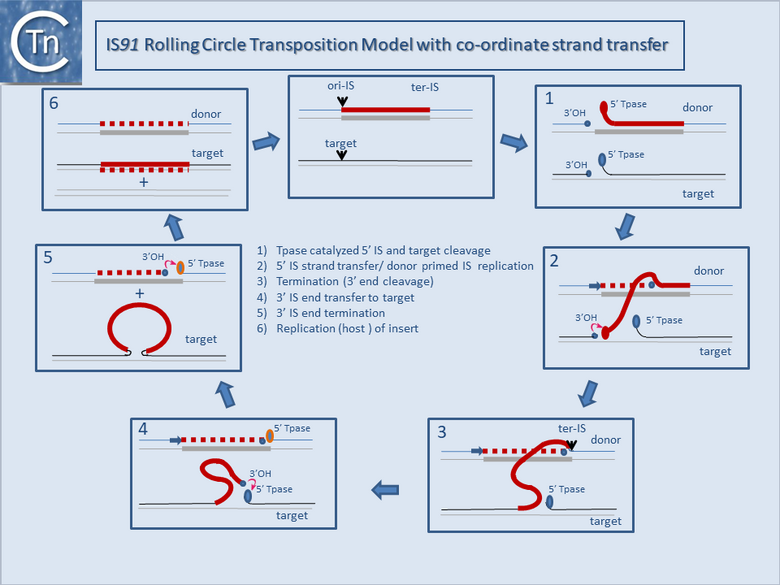
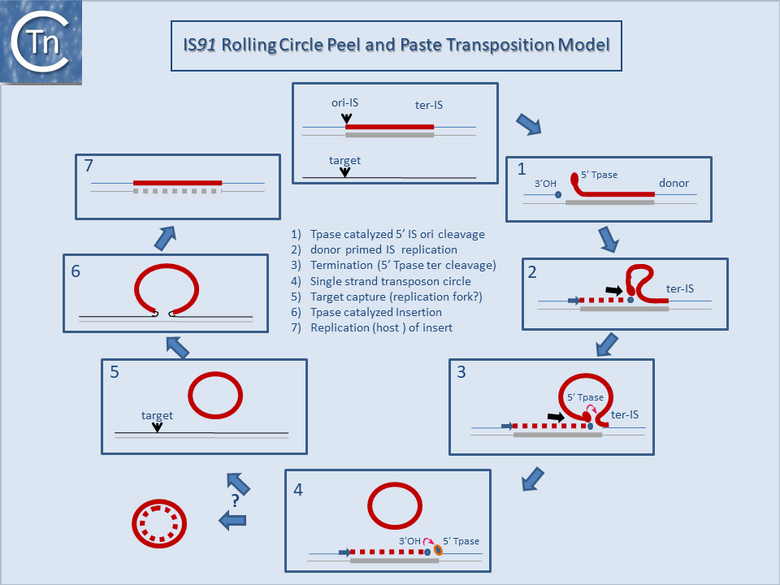
It is thought that these genes are transmitted during the rolling circle type of transposition mechanism postulated to occur in IS91 transposition (Fig.12.2). This involves an initiation event at one IS end, polarized transfer of the IS strand into a target molecule, and termination at the second end[6]. However, the de la Cruz lab has identified circular IS91 forms, and it seems possible that these are transposition intermediates. An alternative model involving transposon circle formation is shown in Fig.12.3. This model is attractive since it would liberate the transposon circle intermediate to locating a target site following circle formation, rather than requiring target engagement during the replicative transposition process (Fig.12.2).
Flanking gene acquisition is thought to occur when the termination mechanism fails and rolling circle transposition extends into neighboring DNA, where it may encounter a second surrogate end [7]. This type of mobile element may prove to play an important role in the assembly and transmission of multiple antibiotic resistance[8].
Bibliography
- ↑ Diaz-Aroca E, de la Cruz F, Zabala JC, Ortiz JM . Characterization of the new insertion sequence IS91 from an alpha-hemolysin plasmid of Escherichia coli. - Mol Gen Genet: 1984, 193(3);493-9 [PubMed:6323920] [DOI]
- ↑ Toleman MA, Bennett PM, Walsh TR . ISCR elements: novel gene-capturing systems of the 21st century? - Microbiol Mol Biol Rev: 2006 Jun, 70(2);296-316 [PubMed:16760305] [DOI]
- ↑ Toleman MA, Walsh TR . ISCR elements are key players in IncA/C plasmid evolution. - Antimicrob Agents Chemother: 2010 Aug, 54(8);3534; author reply 3534 [PubMed:20634542] [DOI]
- ↑ Toleman MA, Bennett PM, Walsh TR . Common regions e.g. orf513 and antibiotic resistance: IS91-like elements evolving complex class 1 integrons. - J Antimicrob Chemother: 2006 Jul, 58(1);1-6 [PubMed:16751201] [DOI]
- ↑ Chandler M, de la Cruz F, Dyda F, Hickman AB, Moncalian G, Ton-Hoang B . Breaking and joining single-stranded DNA: the HUH endonuclease superfamily. - Nat Rev Microbiol: 2013 Aug, 11(8);525-38 [PubMed:23832240] [DOI]
- ↑ Garcillán-Barcia MP, de la Cruz F . Distribution of IS91 family insertion sequences in bacterial genomes: evolutionary implications. - FEMS Microbiol Ecol: 2002 Nov 1, 42(2);303-13 [PubMed:19709290] [DOI]
- ↑ De La Cruz F, Garcillán-Barcia MP, Bernales I, Mendiola MV. IS91 Rolling-Circle Transposition. In: Craig NL, Lambowitz AM, Craigie R, Gellert M, editors. Mobile DNA II. American Society of Microbiology; 2002. p. 891–904.
- ↑ Toleman MA, Walsh TR . Combinatorial events of insertion sequences and ICE in Gram-negative bacteria. - FEMS Microbiol Rev: 2011 Sep, 35(5);912-35 [PubMed:21729108] [DOI]