Mobile element type:
Transposon
Name:
TnXc4.2
Synonyms:
Tn7212
Accession:
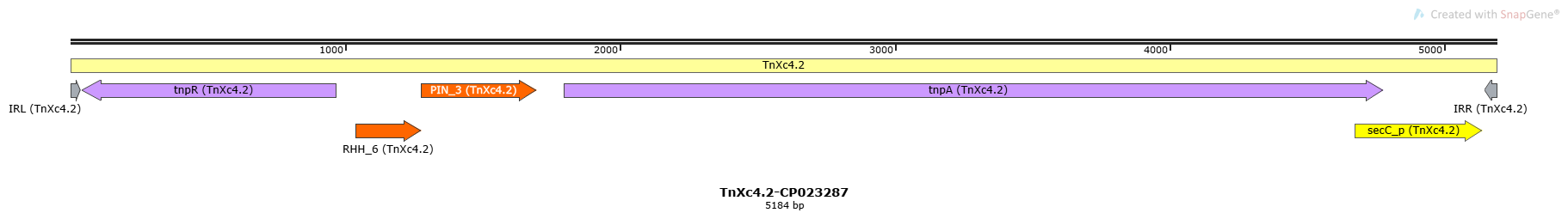
TnXc4.2-CP023287
Family:
Tn3
Group:
Tn3
First isolate:
Yes
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Xanthomonas citri pv. citri 03-1638-1-1
Date of Isolation:
2018
Country:
Argentina
Molecular Source:
plasmid pP2
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGTTCGGG GAGCAATGGA ACAGGGAAGT CAGTTAAG |
| IRR | 5147-5184 | - | 38 | GGGGTTCGGG GAGCAATGGA ACAGGGAAGT CAGTTAAG |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tnpR | TnXc4.2 | 42-965 | - | Accessory Gene | Resolvase |
| RHH_6 | TnXc4.2 | 1039-1278 | + | Passenger Gene | Antitoxin |
| PIN_3 | TnXc4.2 | 1278-1697 | + | Passenger Gene | Toxin |
| tnpA | TnXc4.2 | 1795-4779 | + | Transposase | |
| secC_p | TnXc4.2 | 4673-5140 | + | Passenger Gene | Plant Pathogenicity |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | TnXc4.2 | 307 | 42-965 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MKIGYARVST REQNPALQVD SLKAAGCERI YQDVASGAKT ARPALDELLG QLRGGDVLVI WKLDRMGRSL KHLVELVGSL MERKVGLLSL NDPIDTTSAQ GRFVFNLFAT LAEFERELIR ERTQAGLTAA RARGRVGGRP KGLSPQAEAT ALAAETLYRE RKLSVAAIAQ KLHLSKSTLY SYLRHRGVEI GPYKQSAQSP INVSVVESGN ADQPKVATIL LTLRIENNSK FVRGKKRSIE HVEFFDLAQY DATRRPNGEY ELKVPYDTDE ELDEAVNDLL TDIACGADDR HCFSESHARM EGTDRHW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | RHH_6 | RHH_6 | TnXc4.2 | 79 | 1039-1278 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antitoxin | ||||||||||
| Sequence Family: | RHH_6 (Pfam:PF16762) | ||||||||||
| Protein Sequence: | MNTVRWNIAV SPDVDQSVRM FIAAQGGGRK GDLSRFIEDA VRAYLFERAV EQAKAATVGM GETELNDLID EAVQWAREH |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | PIN_3 | PIN_3 | TnXc4.2 | 139 | 1278-1697 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Toxin | ||||||||||
| Sequence Family: | PIN_3 (Pfam:PF13470) | ||||||||||
| Target: | single stranded RNA | ||||||||||
| Comment: | tRNA(fMet)-specific endonuclease | ||||||||||
| Protein Sequence: | MRVVLDTNVL LAALISSHSP PDIIYRGWLA ARFELVTGTA QLDELRRVSR YPKIKAILPA HRVGTMINNM QRAVVLHVLP PLPDRIEVND PNDAFLLAMA LASEADYLVT GDRRAGLLQR GSIGRTRIVT PVTFCAEAL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | TnXc4.2 | 994 | 1795-4779 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | missing 135 bp at 3' (C-ter) end | ||||||||||
| Protein Sequence: | LDDDDREWIA TKRRDSSRLG YALQLTTARF LGTFLEDPTA VPSPVLHTLS SQLGIADPSD CVIDYRTTRQ RWQHTSEIRT RYGYREFTGT GVQFRLGRWL CALCWTGTDR PSALFDYANG WLVGHKVLLP GVTLLERFIA *IRSRMESRL WRLLVHGVTP EQRQRLDDLL KLVEGSRQSW LDRLRKGPVR VSAPALVAAL LRIETVRGLG IKLPGTHVPP SRIAALARFA STAKVSAVAR LPEVRRIATL VAFVHCLEAS AQDDAIDVLD LLLRELFTKA EKEDRKVRQR SLKDLDRAAS TLAEACRMLL DPALPDGELR ERVYAAIGHD ELAQALNEVR GLVRPPNDVF YTELEARKAT VSRFLPALLR VIRFDANPAA QPLAQALQWL HEKPDHDPPT AIVGKAWQRH VVQDDGRINA TAYSFCALDK LRSAIRRRDV FISPSWRYAD PRAGLLAGAE WEASRPIVCR SLSLSAQPEA TLSELTRELD ETYRRVAARL PQNDAVRFEN VGDKTELVLS PLEALEEPPS LIALRNEIKA RMPRVDLPEI LLEVAGRTGC MEAFTHLTER TARAADLTTS LCAVLMAEAC NTGPEPLVRP DTPALKRDRL MWVDQNYVRD DTLTACNAVL VAAQSRIALA RTWGGGDVAS ADGMRFVVPV RTIHAGPNPK YFNRGRGVTW YNLLSDQRTG LNAITVPGTL RDSLILLAVV LEQQTELQPT QIMTDTGAYS DLVFGLFRLS NYRFCPRLAD VGGTRFWRVD PDADYGDLNA LARQRVNLDR ITPHWDDVLR LVGSLKLGLV PAMGIMRTLQ VDERPTSLAQ AIAEIGRIDK TIHTLNFIDD EARRRATLLQ LNLGEGRHSL AREVFHGKRG ELFQRYREGQ EDQLSALGLV VNMIVLWNTL YMDAVLTQLR SEGYPVKPED EARLSPFGHE HINMLGRYSF SVPEAVARGE PSRRNSWLGH AALEHDCIRI VPHSGERRCL PSRATRRCGW GRTR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | secC_p | SecC_p | TnXc4.2 | 155 | 4673-5140 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Plant Pathogenicity | ||||||||||
| Comment: | C-terminal segment | ||||||||||
| Protein Sequence: | MRHWNTIASE LFRTLEKDDV YLPVLLEDAD GAVHGNDWAR GFMRGIQLRP NSWQELIGSE EFGGPMLPIM ILTHEHDPDP AMRPPEIAPD KRDELLQSLI AGLTHIYRYF ASHRQLATQG SLRRQGPKIG RNDQCPCGSG RKYKHCCATS APTFH |
||||||||||
1. Gochez AM, Huguet-Tapia JC, Minsavage GV, Shantaraj D, Jalan N, Strauss A, Lahaye T, Wang N, Canteros BI, Jones JB, Potnis N.
Pacbio sequencing of copper-tolerant Xanthomonas citri reveals presence of a chimeric plasmid structure and provides insights into reassortment and shuffling of transcription activator-like effectors among X. citri strains.. BMC Genomics. 2018 Jan 4;19(1):16. doi: 10.1186/s12864-017-4408-9. PubMed ID:
29301493
.