Mobile element type:
Transposon
Name:
TnPfu1.1
Synonyms:
Tn7221
Accession:
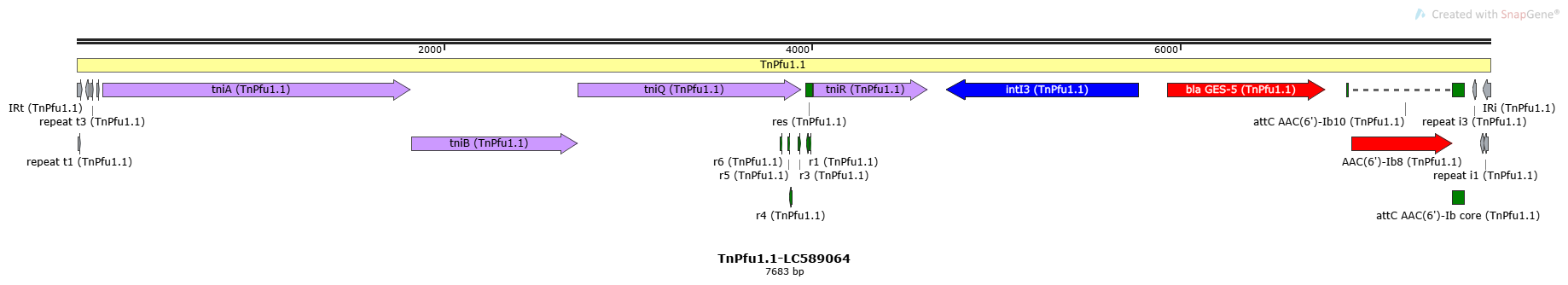
TnPfu1.1-LC589064
Family:
Tn402
Group:
TnPfu1
First isolate:
Yes
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Pseudomonas aeruginosa IPM3H3
Date of Isolation:
2020
Country:
Japan
Molecular Source:
plasmid pIPM3H3-GES5
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRt | 1-33 | + | 33 | TGTCATTTTC AGAAGACGAC TGCACCAATT GAT |
| repeat t1 | 9-27 | + | 19 | TCAGAAGACG ACTGCACCA |
| repeat t2 | 49-67 | - | 19 | TCAGGAGCTG GCTGCACAA |
| repeat t3 | 78-97 | + | 20 | TCAGAAGTGA TCTGCACCAA |
| repeat t4 | 110-128 | + | 19 | TCAATACTCG TGTGCACCA |
| repeat i3 | 7594-7613 | - | 20 | TCATGCAATG GCGGGCAGCA |
| repeat i2 | 7634-7652 | - | 19 | TCAGAAGCCG ACTGCACTA |
| IRi | 7651-7683 | - | 33 | TGTCGTTTTC AGAAGGCGGC TGCACTGAAC GTC |
| repeat i1 | 7656-7675 | - | 20 | TCAGAAGGCG GCTGCACTGA |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| r6 | TnPfu1.1 | 3824-3837 | + | 14 | TTGGAACGGT TGCG |
| r5 | TnPfu1.1 | 3870-3883 | + | 14 | AATTTCTGTC ACAT |
| r4 | TnPfu1.1 | 3875-3888 | - | 14 | AGGTAATGTG ACAG |
| r3 | TnPfu1.1 | 3926-3939 | + | 14 | GGGTTAAGTG ACAA |
| res | TnPfu1.1 | 3967-4000 | + | 34 | ACACTGTCAC ATAATCGAAC GTATACGTGA CAGG |
| r2 | TnPfu1.1 | 3970-3983 | - | 14 | CGATTATGTG ACAG |
| r1 | TnPfu1.1 | 3986-3999 | + | 14 | CGTATACGTG ACAG |
| attC AAC(6')-Ib10 | TnPfu1.1 | 6908-6914 , 7482-7547 | + | 73 | GTTAGGCGCC TAACCCTTCC ATCGAGGGGG ACGTCCAAGG GCTGGCGCCC TTGGCCGCCC CTCATGTCAA ACG |
| attC AAC(6')-Ib core | TnPfu1.1 | 7482-7547 | + | 66 | GCCTAACCCT TCCATCGAGG GGGACGTCCA AGGGCTGGCG CCCTTGGCCG CCCCTCATGT CAAACG |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tniA | TnPfu1.1 | 142-1821 | + | Transposase | |
| tniB | TnPfu1.1 | 1824-2732 | + | Accessory Gene | |
| tniQ | TnPfu1.1 | 2729-3946 | + | Accessory Gene | Target Site Selection |
| tniR | TnPfu1.1 | 4008-4631 | + | Accessory Gene | Resolvase |
| intI3 | TnPfu1.1 | 4733-5773 | - | Integron Integrase | Class 3 |
| bla GES-5 (ARO:3002334) | TnPfu1.1 | 5932-6795 | + | Passenger Gene | Antibiotic Resistance |
| AAC(6')-Ib8 (ARO:3002579) | TnPfu1.1 | 6936-7487 | + | Passenger Gene | Antibiotic Resistance |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA | TniA | TnPfu1.1 | 559 | 142-1821 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | can be extended upstream by 12 amino acids| identical to tniA (Tn1721 and In2)| 25% amino acid sequence identity to TnsB from Tn7 | ||||||||||
| Protein Sequence: | MATDTPRIPE QGVATLPDEA WERARRRAEI ISPLAQSETV GHEAADMAAQ ALGLSRRQVY VLIRRARQGS GLVTDLVPGQ SGGGKGKGRL PEPVERVIHE LLQKRFLTKQ KRSLAAFHRE VTQVCKAQKL RVPARNTVAL RIASLDPRKV IRRREGQDAA RDLQGVGGEP PAVTAPLEQV QIDHTVIDLI VVDDRDRQPI GRPYLTLAID VFTRCVLGMV VTLEAPSAVS VGLCLVHVAC DKRPWLEGLN VEMDWQMSGK PLLLYLDNAA EFKSEALRRG CEQHGIRLDY RPLGQPHYGG IVERIIGTAM QMIHDELPGT TFSNPDQRGD YDSENKAALT LRELERWLTL AVGTYHGSVH NGLLQPPAAR WAEAVARVGV PAVVTRATSF LVDFLPILRR TLTRTGFVID HIHYYADALK PWIARRERWP SFLIRRDPRD ISRIWVLEPE GQHYLEIPYR TLSHPAVTLW EQRQALAKLR QQGREQVDES ALFRMIGQMR EIVTSAQKAT RKARRDADRR QHLKTSARPD KPVPPDTDIA DPQADNLPPA KPFDQIEEW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB | TniB | TnPfu1.1 | 302 | 1824-2732 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Sequence Family: | ATP binding protein? | ||||||||||
| Comment: | similar function to Tn7 tnsC and MuB | ||||||||||
| Protein Sequence: | MDEYPIIDLS HLLPAAQGLA RLPADERVQR LRADRWIGYP RAVEALNRLE ALYAWPNKQR MPNLLLVGPT NNGKSMIVEK FRRTHPASAD ADQEHIPVLV VQMPSEPSVI RFYVALLAAM GAPLRPRPRL PEMEQLALAL LRKVGVRMLV IDELHNVLAG NSVNRREFLN LLRFLGNELR IPLVGVGTRD AYLAIRSDDQ LENRFEPMLL PVWEANDDCC SLLASFAASL PLRRPSSIAT LDMARYLLTR SEGTIGELAH LLMAAAVAAV ESGEEAINHR TLSMADYTGP SERRRQFERE LM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniQ | TniQ | TnPfu1.1 | 405 | 2729-3946 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Target Site Selection | ||||||||||
| Comment: | similar function to Tn7 tnsC and MuB | ||||||||||
| Protein Sequence: | VKPAPRWPLH PAPKEGEALS SWLNRVALCY HMELPELLEH DLGHSQVDDL DTAPPLSLLA MLSQRSGIDP DRLRSMSFAG WVPWLLDSLD DQIPDALETY AFQLSVLLPK LHRKKRSITR WRAWLPSQPI HCACPLCLSD PDNQAVLLAW KLPLMLSCPL HGCWLESYWG VPGRFLGWEN ADAEPRTASD AIAAMDQRTW QALTTGHVEL PCRRIHAGLW FRLLRTLLDE LNTPLSACGT CAGYPRQVWE GCGHPLRAGQ SLWRPYETLN PVVRLQMLEA AATAISLIEV RDISPPGEQA KLFWSEPQTG FTSGLPTKAP KPEPINHWQR AVQAIDEAII EARHNPETAR SLFALASYGR RDPASLERLR ATFVKEGIPP EFLSHYLPDA PFACLKQNDG LSDKF |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniR | TniR | TnPfu1.1 | 207 | 4008-4631 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Comment: | also calledtniR | ||||||||||
| Protein Sequence: | MLIGYMRVSK ADGSQSTDLQ RDALIVAGVS PAHLYEDMAS GRRDDRPGLA ACLKALREGD TLIVWKLDRL GRDLRHLINT VHDLTARSVG LKVLAGHGAA VDTTTAAGKL VFGIFAALAE FERELISERT VAGLVSARAR GRKGGRPFKM TATKLRLAMA SMGQPETKVG NLCEELGITR QTLYRHVSPK GELRPDGVKL LSRGSAA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI3 | IntI3 | TnPfu1.1 | 346 | 4733-5773 | - |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 3 | ||||||||||
| Sequence Family: | Class 3 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MNRYNGSAKP DWVPPRSIKL LDQVRERVRY LHYSLQTEKA YVYWAKAFVL WTARSHGGFR HPREMGQAEV EGFLTMLATE KQVAPATHRQ ALNALLFLYR QVLGMELPWM QQIGRPPERK RIPVVLTVQE VQTLLSHMAG TEALLAALLY GSGLRLREAL GLRVKDVDFD RHAIIVRSGK GDKDRVVMLP RALVPRLRAQ LIQVRAVWGQ DRATGRGGVY LPHALERKYP RAGESWAWFW VFPSAKLSVD PQTGVERRHH LFEERLNRQL KKAVVQAGIA KHVSVHTLRH SFATHLLQAG TDIRTVQELL GHSDVSTTMI YTHVLKVAAG GTSSPLDALA LHLSPG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | bla GES-5 (ARO:3002334) | Bla GES-5 | TnPfu1.1 | 287 | 5932-6795 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | GES beta-lactamase (ARO:3000066) | ||||||||||
| Target: | cephalosporin (ARO:0000032)||penam (ARO:3000008)||carbapenem (ARO:0000020) | ||||||||||
| Protein Sequence: | MRFIHALLLA GIAHSAYASE KLTFKTDLEK LEREKAAQIG VAIVDPQGEI VAGHRMAQRF AMCSTFKFPL AALVFERIDS GTERGDRKLS YGPDMIVEWS PATERFLASG HMTVLEAAQA AVQLSDNGAT NLLLREIGGP AAMTQYFRKI GDSVSRLDRK EPEMSDNTPG DLRDTTTPIA MARTVAKVLY GGALTSTSTH TIERWLIGNQ TGDATLRAGF PKDWVVGEKT GTCANGGRND IGFFKAQERD YAVAVYTTAP KLSAVERDEL VASVGQVITQ LILSTDK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | AAC(6')-Ib8 (ARO:3002579) | AAC(6')-Ib8 | TnPfu1.1 | 183 | 6936-7487 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | AAC(6') (ARO:3000345) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002579 missing 43 aa N-ter (bitscore: 379) | ||||||||||
| Protein Sequence: | TNSTDSVTLR LMTEHDLAML YEWLNRSHIV EWWGGEEARP TLADVQEQYL PSVLAQESVT PYIAMLNGEP IGYAQSYVAL GSGDGWWEEE TDPGVRGIDQ SLANASQLGK GLGTKLVRAL VELLFNDPEV TKIQTDPSPS NLRAIRCYEK AGFERQGTVT TPDGPAVYMV QTRQAFERTR SDA |
||||||||||