Mobile element type:
Transposon
Name:
TnAs2
Synonyms:
Tn7144
Accession:
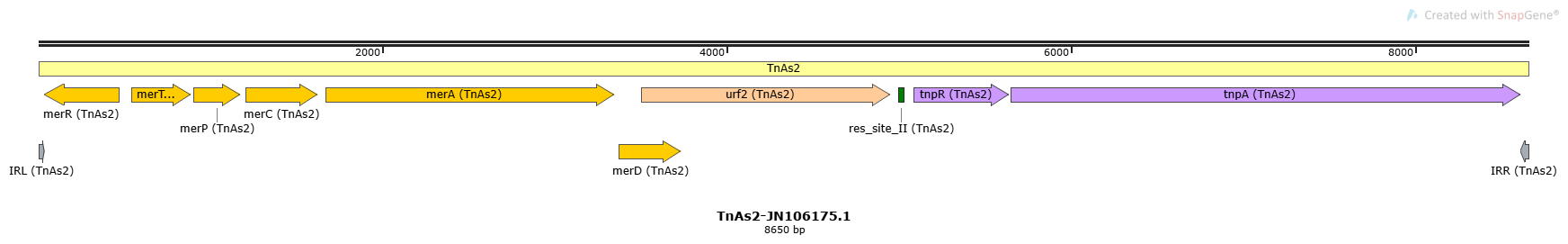
TnAs2-JN106175.1
Family:
Tn3
Group:
Tn21
First isolate:
Yes
Partial:
ND
Evidence of transposition:
Yes
Host Organism:
Uncultured bacterium
Date of Isolation:
2011
Country:
Molecular Source:
plasmidpAKD34
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGTCGTCT CAGAAAACGG AAAATAAAGC ACGCTAAG |
| IRR | 8610-8650 | - | 41 | GGGGTCGCCT CAGAAAACGG AAAATAAAGC ACGCTAAGCC G |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| res_site_II | TnAs2 | 4996-5030 | + | 35 | CGTCAGAATA GAGTCGATTG TGTTATTTAT TGACA |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| merR | TnAs2 | 34-468 | - | Passenger Gene | Heavy Metal Resistance |
| merT | TnAs2 | 540-890 | + | Passenger Gene | Heavy Metal Resistance |
| merP | TnAs2 | 903-1178 | + | Passenger Gene | Heavy Metal Resistance |
| merC | TnAs2 | 1206-1628 | + | Passenger Gene | Heavy Metal Resistance |
| merA | TnAs2 | 1668-3353 | + | Passenger Gene | Heavy Metal Resistance |
| merD | TnAs2 | 3371-3736 | + | Passenger Gene | Heavy Metal Resistance |
| urf2 | TnAs2 | 3501-4955 | + | Passenger Gene | Other |
| tnpR | TnAs2 | 5085-5645 | + | Accessory Gene | Resolvase |
| tnpA | TnAs2 | 5648-8617 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merR | MerR | TnAs2 | 144 | 34-468 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MENNLENLTI GVFAKAAGVN VETIRFYQRK GLLPEPDKPY GSIRRYGAAD VTRVRFVKSA QRLGFSLDEI ADLLRLDDGT HCEEASSLAE HKLQNVREKM ADLARMETVL SELVCACHAR KGNVSCPLIA SLQGGTSLAG SAMP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merT | MerT | TnAs2 | 116 | 540-890 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSEPQTGRGA LFAGGLAAIL ASACCLGPLV LIALGFSGAW IGNLTVLEPY RPIFIGAALV ALFFAWRRIY RPAQACRPGE VCAIPQVRAT YKLIFWVVAV LVLVALGFPY VMPFFY |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merP | MerP | TnAs2 | 91 | 903-1178 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MKKLFASLAL AAVVAPVWAA TQTVTLSVPG MTCSACPITV KKAISKVDGV SKVAVTFETR EAVVTFDDAK TSVQKLTKAT GDAGYPSSVK Q |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merC | MerC | TnAs2 | 140 | 1206-1628 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MGLITRIADK AGALGSVVSA MGCAACFPAI ASLGAAIGLG FLQEYEGLFI FTLLPLFAVV ALLANALGWL SHRQWHRSLL GMIGPAIVFA GTVWLLGNWW TARLVYTGLA LMIGVSIWDL VSPANRRCGP DGCELPAKHG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merA | MerA | TnAs2 | 561 | 1668-3353 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MTTLKITGMT CDSCAVHVKE ALEKVPGVQS ANVSYTKGSA KLAVETGTSP DALTAAVAGL GYRATLADAL VPPVGGGLLV KMREWLGSGD KAGDDGGGLH IAVIGSGGAA MAAALKAVEQ GAHVTLIERG TIGGTCVNIG CVPSKIMIRA AHIAHLRRES PFDGGMPPTP PTILRERLLA QQQARVDELR HAKYEGILDD NPAISVLHGE ARFKDGHSLT VQLNGGGERV VTFDRCLIAT GASPAVPPIP GLKDTPYWTS TEALVSDTIP ERLAVIGSSV VALELAQAFA RLGSQVTILA RSTLFFREDP AIGEAITAAF RAEGIEVLEH TQASQVAHEG GEFVLTTAHG ELRADKLLVA TGRSPNTRSL ALDAAGVALN LQGAIVIDVG MRTSTPDIYA AGDCTDQPQF VYVAAAAGTR AAINMTGGDA ALNLAAMPAV VFTDPQVATV GYSEAEAQHD GIETDSRTLT LDNVPRALAN FDTRGFIKLV IEEGSGRLIG VQAVAPEAGE LIQTAVLAIR NRMTVQELAD QLFPYLTMVE GLKLAAQTFN KDVKQLSCCA G |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merD | MerD | TnAs2 | 121 | 3371-3736 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MNAYTVSRLA LNAGVSVHIV RDYLLRGLLR PVACTPGGYG LFDDAALQRL CFVRAAFEAG IGLDALARLC RALDTADGDE AAAQLAVLRQ FVERRREALA DLEVQLATMP TELVQHAESL P |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | urf2 | Urf2 | TnAs2 | 484 | 3501-4955 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | EAL (Pfam:PF00563)||DUF3330 (Pfam:PF11809) | ||||||||||
| Comment: | similar to urfM from E.coli | ||||||||||
| Protein Sequence: | MRPCNGCASC GLLSKRVSVS TRWRGCAGRW IPRTATKRPR SLPCCASSSN VGAKHWPIWR CSWPPCRPSW YSMRRVCHEQ PRALADRDTQ AVHRLPVGRV GRAHLSLPFA DSRCGARGDD GRRVHRGALG YCSHHADRLV CPVRDAAVAG LQGKIVSTSL PARWTATELA QAVVRGQFEL HYQPIVDLRS DQIVGAEALL RWRHPQLGLL PPGQFLPVVE SSGLMPEIGA WVLGTACRQM REWRMLAWQP FRLAVNVSAS QVGPDFDGWV KGVLADAELP AEYLEIELTE SVAFGNPAIF PALEALRQIG VRFAADDFGT GYSCLQHLKC CPITKLKIDQ SFVAGLADDH RDRTIVHTVI QLAHGLGMDV VAEGVETPTS LALLRQAECD TGQGFLFAKP VPAAAFAAFV SQWRGATMNA NDPTATSCCV CCKEIPLDAA FTPEGAEYVE RFCGLECYQR FQARASNATE TSAELNACGS PPSD |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | TnAs2 | 186 | 5085-5645 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Function: | resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MQGQRIGYVR VSSFDQNPER QLEHVEVGKV FTDKASGKDT QRPELDSLLA FVREGDTVVV HSMDRLARNL DDLRRLVQKL TKRGVRIEFV KESLTFTGED SPMANLMLSV MGAFAEFERA LIRERQREGI ALAKQRGAYR GRKKALSPEQ VADLRQRAAA GEQKAKLARE FGVSRETLYQ YLRTDQ |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | TnAs2 | 989 | 5648-8617 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MPRRSILSAA ERESLLALPD SKDDLIRHYT FSDSDLSIIR QRRGPANRLG FAVQLCYLRF PGIILGVDQP PFLPLLKLVA DQLKVGIENW DEYGQREQTR REHLVELQAL FGFQPFTMSH YRQAVHTLTE LAMQTDKGIV LASALIEHLR RQSVILPALN AVERASAEAI TRANRRIYDA LAEPLSDAHR RRLDDLLKRR DNGKTTWLAW LRQSPVKPNS RHMLEHIERL KAWQTLDLPS GIERLVHQNR LLKIAREGGQ MTPADLAKFE PQRRYATLVA LAIEGMATVT DEIIDLHDRI LGKLFNAAKN KHQQQFQASG KAINAKVRLY GRIGQALIDA KQSGRDPFAA IEAVMSWDAF AESVTEAQKL AQPDDFDFLH RIGESYATLR RYAPEFLAVL KLRAAPAAKN VLDAIEVLRG MNTDNARKVP ADAPTDFIKP RWQKLVMTDA GIDRRYYELC ALSELKNSLR SGDIWVQGSR QFKDFEDYLV PPAKFASLKQ SSELPLAVAT DCDQYLHERL TLLETQLATV NRMAAANNLP DAIITESGLK ITPLDAAVPD TAQALIDQTA MILPHVKITE LLLEVDEWTG FTRHFTHLKS GDLAKDKNLL LTTILADAIN LGLTKMAESC PGTTYAKLAW LQAWHTRDET YSTALAELVN AQFQHPFAGH WGDGTTSSSD GQNFRTGSKA ESTGHINPKY GSSPGRTFYT HISDQYAPFH TKVVNVGVRD STYVLDGLLY HESDLRIEEH YTDTAGFTDH VFALMHLLGF RFAPRIRDLG DTKLFIPKGE TNYDALKPMI SSDKLNIKAI RAHWDEILRL ATSIKQGTVT ASLMLRKLGS YPRQNGLAVA LRELGRIERT LFILDWLQSV ELRRRVHAGL NKGEARNALA RAVFFNRLGE IRDRSFEQQR YRASGLNLVT AAVVLWNTVY LERAAHALRG NGHAVDDALL QYLSPLGWEH INLTGDYLWR SSAKIGAGKF RPLRPLQPA |
||||||||||
1. Sen D, Van der Auwera GA, Rogers LM, Thomas CM, Brown CJ, Top EM.
Broad-host-range plasmids from agricultural soils have IncP-1 backbones with diverse accessory genes.. Appl Environ Microbiol. 2011 Nov;77(22):7975-83. doi: 10.1128/AEM.05439-11. Epub 2011 Sep 23. PubMed ID:
21948829
.