Mobile element type:
Transposon
Name:
TnAmu1
Synonyms:
Tn7138
Accession:
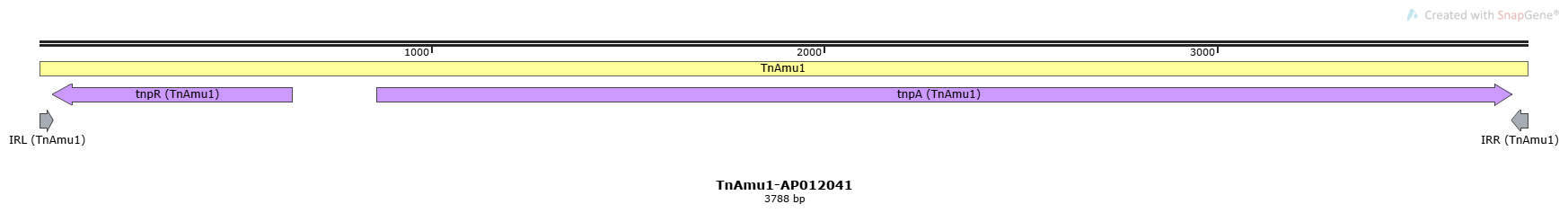
TnAmu1-AP012041
Family:
Tn3
Group:
Tn163
First isolate:
Yes
Partial:
ND
Evidence of transposition:
Yes
Host Organism:
Acidiphilium multivorum AIU301
Date of Isolation:
2010
Country:
Molecular Source:
plasmid pACMV6
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGTCACCG CACGATCTGT TCAAAATCGT ACGCTAAG |
| IRR | 3751-3788 | - | 38 | GGGGTCACCG CACGATCTGT TCAAAATCGT ACGCTAAG |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tnpR | TnAmu1 | 34-645 | - | Accessory Gene | Resolvase |
| tnpA | TnAmu1 | 861-3755 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | TnAmu1 | 203 | 34-645 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MKGGHTLQFR MAHTLIGYAR CSTDRQDLAA QRAELEQLGV KPDRIYTDHG LTGTNRARPG LAQALAAVRK GDTLVVPKLD RLARSVPDAR AIADELITRG VKLALGAAVY DPTDPMGKMF FNILATFAEF EADLIRLRTR EGMAIARAKG KLRGKRPTLS ERQQKELRRM YDTGNYSITD LADLFSVSRP TIYRTLARQP ANA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | TnAmu1 | 964 | 861-3755 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MSRHYVLDAA DLAMVRVRRR ASNRLGFAVQ LCVLRFPGRT LDPSEYPPAP ALAFVAEQIG VDPLLFAEYA RRAETRREHL VELQKFTGLR SFGLDDWRAC LHAGADTAWA TDRGEPIIQA MLVHLRESRV LLPSATVLER IGLAARVRAR KKTFEVLAAG MADAERSALT ELLTVDPELR RSRFAWLRDY SESPAPSNIV SLLDRLEYVR AMGIDPARAG RIHAARLVRL TDEGALMTVQ HIADLEPARR TAILIAQIAS LETRLTDATL AMFEKYVGTL FSRARNRDER RFQATRRDVA KALLLFRRTI AALRQAKEAG EDGITAIERE IGMNQLEDVL PIIGAVADVA DQDILVTAAE RYTVLRRFSP RFLAAFDFRS NMPNDPVLAA IELLRALNRG AIRSLPKRPP STFLPPQWRK LIFAGGTADR RLYETAVLAV LRDKLRGSNI WVAGSRDYQA FETYLLPAGA GAATGIDGEA DPDRYVASRA EMLRERLTFV AARAAQGDLD GVEIEDGKLF IARTPPTVPD AARDLALRLN SMLPRVRITE VLSEVDAWTG FTDRFVHLRT GNPVTDKAAL LAAVLADGTN LGLARMADAS RGLSYHHLVN VAQWHISDDN YVAARAAIIN AHHAHPMAAI WDDGTSSSSD GQYFRAAGRA GAGGSVNAKY GIEPGAVIYT HVSGQYGPFH TRVISATMSE APYVLDGLLH HVHQTDLRIA EHYTDTAGAT DHVFGLCHLL GYRFAPRIKD LRDRKLYTIE KPGTWPLLVP LIGDAVETTA ILGQWPELMR LKASINAGTV LPSVILRKLA AAGGGNTLSR ALRAVGRIER TLFTLQWLSD PALRQRSHAG LNKGEASNAL RRAVFFHRQG EIRDRTFENQ SFRASGLSLI TAAIVHWNTI YLDQAVQHLR AQGTAVPDDL LAHVAPLGWE HIALTGDYVW NPANPNASFR PLRDVRAPFI PRAA |
||||||||||