Mobile element type:
Transposon
Name:
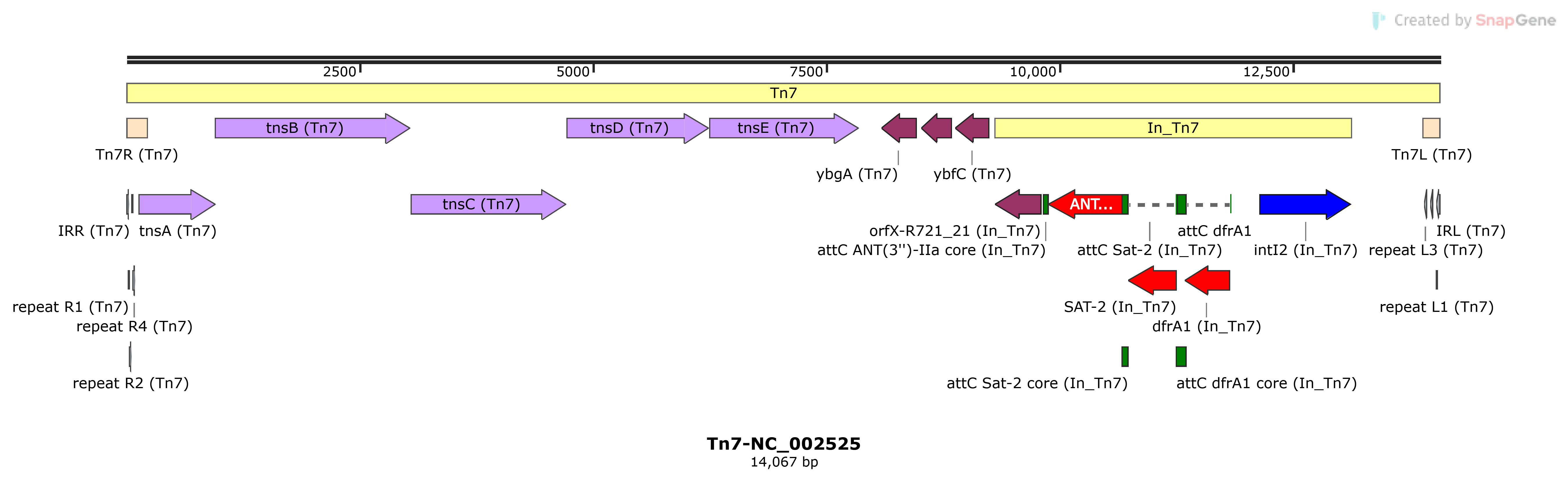
Tn7
Synonyms:
Accession:
Tn7-NC_002525
Family:
Tn7
Group:
First isolate:
Yes
Partial:
ND
Evidence of transposition:
Yes
Host Organism:
Escherichia coli
Date of Isolation:
1972
Country:
Molecular Source:
plasmid R721
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| Tn7R | 1-225 | + | 225 | TGTGGGCGGA CAATAAAGTC TTAAACTGAA CAAAATAGAT CTAAACTATG ACAATAAAGT CTTAAACTAG ACAGAATAGT TGTAAACTGA AATCAGTCCA GTTATGCTGT GAAAAAGCAT ACTGGACTTT TGTTATGGCT AAAGCAAACT CTTCATTTTC TGAAGTGCAA ATTGCCCGTC GTATTAAAGA GGGGCGTGGC CAAGGGCATG GTAAAGACTA TATTC |
| IRR | 1-30 | + | 30 | TGTGGGCGGA CAATAAAGTC TTAAACTGAA |
| repeat R1 | 9-30 | + | 22 | GACAATAAAG TCTTAAACTG AA |
| repeat R2 | 29-50 | + | 22 | AACAAAATAG ATCTAAACTA TG |
| repeat R3 | 50-71 | + | 22 | GACAATAAAG TCTTAAACTA GA |
| repeat R4 | 70-91 | + | 22 | GACAGAATAG TTGTAAACTG AA |
| Tn7L | 13902-14067 | + | 166 | AACCAGATAA GTGAAATCTA GTTCCAAACT ATTTTGTCAT TTTTAATTTT CGTATTAGCT TACGACGCTA CACCCAGTTC CCATCTATTT TGTCACTCTT CCCTAAATAA TCCTTAAAAA CTCCATTTCC ACCCCTCCCA GTTCCCAACT ATTTTGTCCG CCCACA |
| repeat L3 | 13918-13940 | - | 23 | TGACAAAATA GTTTGGAACT AGA |
| repeat L2 | 13974-13996 | - | 23 | TGACAAAATA GATGGGAACT GGG |
| repeat L1 | 14038-14060 | - | 23 | GGACAAAATA GTTGGGAACT GGG |
| IRL | 14040-14067 | - | 28 | TGTGGGCGGA CAAAATAGTT GGGAACTG |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| attC ANT(3'')-IIa core | In_Tn7 | 9823-9882 | + | 60 | CTCTAACGCT TGAGTTAAGC CGCGCCGCGA AGCGGCGTCG GCTTGAACGA ATTGTTAGAC |
| attC Sat-2 | In_Tn7 | 10676-10735 , 11260-11265 | + | 66 | GTTTAACTCC TGAATTAAGC CGCGCCGCGA AGCGGTGTCG GCTTGAATGA ATTGTTAGGC GCCTAA |
| attC Sat-2 core | In_Tn7 | 10676-10735 | + | 60 | GTTTAACTCC TGAATTAAGC CGCGCCGCGA AGCGGTGTCG GCTTGAATGA ATTGTTAGGC |
| attC dfrA1 core | In_Tn7 | 11260-11354 | + | 95 | GCCTAACGCC TGGCACAGCG GATCGCAAAC CTGGCGCGGC TTTTGGTACA AAAGGCGTGA CAGGTTTGCG AATCCGTTGC TGCCACTTGT TAACC |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tnsA | Tn7 | 135-956 | + | Transposase | |
| tnsB | Tn7 | 943-3051 | + | Transposase | |
| tnsC | Tn7 | 3048-4715 | + | Accessory Gene | Helper |
| tnsD | Tn7 | 4718-6244 | + | Accessory Gene | Target Site Selection |
| tnsE | Tn7 | 6245-7861 | + | Accessory Gene | Target Site Selection |
| ybgA | Tn7 | 8092-8469 | - | Passenger Gene | Hypothetical |
| ybfB | Tn7 | 8523-8846 | - | Passenger Gene | Hypothetical |
| ybfC | Tn7 | 8879-9250 | - | Passenger Gene | Hypothetical |
| orfX-R721_21 | In_Tn7 | 9311-9808 | - | Passenger Gene | Hypothetical |
| ANT(3'')-IIa (ARO:3004089) | In_Tn7 | 9884-10672 | - | Passenger Gene | Antibiotic Resistance |
| SAT-2 (ARO:3002895) | In_Tn7 | 10730-11254 | - | Passenger Gene | Antibiotic Resistance |
| dfrA1 (ARO:3002854) | In_Tn7 | 11349-11822 | - | Passenger Gene | Antibiotic Resistance |
| intI2 | In_Tn7 | 12154-13131 | + | Integron Integrase | Class 2 |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnsA | TnsA | Tn7 | 273 | 135-956 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase | ||||||||||
| Comment: | resembles a type II restriction enzyme||functions at the 5' end of the transposon||interacts with TnsB||forms double stranded breaks at ends of transposon | ||||||||||
| Protein Sequence: | MAKANSSFSE VQIARRIKEG RGQGHGKDYI PWLTVQEVPS SGRSHRIYSH KTGRVHHLLS DLELAVFLSL EWESSVLDIR EQFPLLPSDT RQIAIDSGIK HPVIRGVDQV MSTDFLVDCK DGPFEQFAIQ VKPAAALQDE RTLEKLELER RYWQQKQIPW FIFTDKEINP VVKENIEWLY SVKTEEVSAE LLAQLSPLAH ILQEKGDENI INVCKQVDIA YDLELGKTLS EIRALTANGF IKFNIYKSFR ANKCADLCIS QVVNMEELRY VAN |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnsB | TnsB | Tn7 | 702 | 943-3051 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | functions at the 3' end of the transposon||interacts with TnsA||also interacts with target site for joining reaction | ||||||||||
| Protein Sequence: | MWQINEVVLF DNDPYRILAI EDGQVVWMQI SADKGVPQAR AELLLMQYLD EGRLVRTDDP YVHLDLEEPS VDSVSFQKRE EDYRKILPII NSKDRFDPKV RSELVEHVVQ EHKVTKATVY KLLRRYWQRG QTPNALIPDY KNSGAPGERR SATGTAKIGR AREYGKGEGT KVTPEIERLF RLTIEKHLLN QKGTKTTVAY RRFVDLFAQY FPRIPQEDYP TLRQFRYFYD REYPKAQRLK SRVKAGVYKK DVRPLSSTAT SQALGPGSRY EIDATIADIY LVDHHDRQKI IGRPTLYIVI DVFSRMITGF YIGFENPSYV VAMQAFVNAC SDKTAICAQH DIEISSSDWP CVGLPDVLLA DRGELMSHQV EALVSSFNVR VESAPPRRGD AKGIVESTFR TLQAEFKSFA PGIVEGSRIK SHGETDYRLD ASLSVFEFTQ IILRTILFRN NHLVMDKYDR DADFPTDLPS IPVQLWQWGM QHRTGSLRAV EQEQLRVALL PRRKVSISSF GVNLWGLYYS GSEILREGWL QRSTDIARPQ HLEAAYDPVL VDTIYLFPQV GSRVFWRCNL TERSRQFKGL SFWEVWDIQA QEKHNKANAK QDELTKRREL EAFIQQTIQK ANKLTPSTTE PKSTRIKQIK TNKKEAVTSE RKKRAEHLKP SSSGDEAKVI PFNAVEADDQ EDYSLPTYVP ELFQDPPEKD ES |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnsC | TnsC | Tn7 | 555 | 3048-4715 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Helper | ||||||||||
| Function: | adaptor protein | ||||||||||
| Comment: | ATP-dependent DNA-binding protein||interacts with double-stranded DNA nonspecifically and the transposase TnsAB and the target site selectors TnsD and E||ATP-hydrolysis promotes correct transposition | ||||||||||
| Protein Sequence: | MSATRIQAVY RDTGVEAYRD NPFIEALPPL QESVNSAASL KSSLQLTSSD LQKSRVIRAH TICRIPDDYF QPLGTHLLLS ERISVMIRGG YVGRNPKTGD LQKHLQNGYE RVQTGELETF RFEEARSTAQ SLLLIGCSGS GKTTSLHRIL ATYPQVIYHR ELNVEQVVYL KIDCSHNGSL KEICLNFFRA LDRALGSNYE RRYGLKRHGI ETMLALMSQI ANAHALGLLV IDEIQHLSRS RSGGSQEMLN FFVTMVNIIG VPVMLIGTPK AREIFEADLR SARRGAGFGA IFWDPIQQTQ RGKPNQEWIA FTDNLWQLQL LQRKDALLSD EVRDVWYELS QGVMDIVVKL FVLAQLRALA LGNERITAGL LRQVYQDELK PVHPMLEALR SGIPERIARY SDLVVPEIDK RLIQLQLDIA AIQEQTPEEK ALQELDTEDQ RHLYLMLKED YDSSLLIPTI KKAFSQNPTM TRQKLLPLVL QWLMEGETVV SELEKPSKSK KVSAIKVVKP SDWDSLPDTD LRYIYSQRQP EKTMHERLKG KGVIVDMASL FKQAG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnsD | TnsD | Tn7 | 508 | 4718-6244 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Target Site Selection | ||||||||||
| Function: | target site selection | ||||||||||
| Comment: | DNA-binding protein||functions with the TnsABC complex toselect specific target sites in the chromosome called attTn7 (this 63-bp site is at the 3' end of the glutamine synthase gene glmS as well as other targets in other chromosomes) | ||||||||||
| Protein Sequence: | MRNFPVPYSN ELIYSTIARA GVYQGIVSPK QLLDEVYGNR KVVATLGLPS HLGVIARHLH QTGRYAVQQL IYEHTLFPLY APFVGKERRD EAIRLMEYQA QGAVHLMLGV AASRVKSDNR FRYCPDCVAL QLNRYGEAFW QRDWYLPALP YCPKHGALVF FDRAVDDHRH QFWALGHTEL LSDYPKDSLS QLTALAAYIA PLLDAPRAQE LSPSLEQWTL FYQRLAQDLG LTKSKHIRHD LVAERVRQTF SDEALEKLDL KLAENKDTCW LKSIFRKHRK AFSYLQHSIV WQALLPKLTV IEALQQASAL TEHSITTRPV SQSVQPNSED LSVKHKDWQQ LVHKYQGIKA ARQSLEGGVL YAWLYRHDRD WLVHWNQQHQ QERLAPAPRV DWNQRDRIAV RQLLRIIKRL DSSLDHPRAT SSWLLKQTPN GTSLAKNLQK LPLVALCLKR YSESVEDYQI RRISQAFIKL KQEDVELRRW RLLRSATLSK ERITEEAQRF LEMVYGEE |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnsE | TnsE | Tn7 | 538 | 6245-7861 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Target Site Selection | ||||||||||
| Function: | target site selection | ||||||||||
| Comment: | DNA-binding protein||functions with the TnsABC complex toselect target sites for transposition similar to TnsD function||TnsE specifically targets sites with orientation bias the same as DNA replication, which leads to insertion into actively conjugating plasmid molecules in recipient cells undergoing lagging strand synthesis | ||||||||||
| Protein Sequence: | MVRLATFNDN VQVVHIGHLF RNSGHKEWRI FVWFNPMQER KWTRFTHLPL LSRAKVVNST TKQINKADRV IEFEASDLQR AKIIDFPNLS SFASVRNKDG AQSSFIYEAE TPYSKTRYHI PQLELARSLF LINSYFCRSC LSSTALQQEF DVQYEVERDH LEIRILPSSS FPKGALEQSA VVQLLVWLFS DQDVMDSYES IFRHYQQNRE IKNGVESWCF SFDPPPMQGW KLHVKGRSSN EDKDYLVEEI VGLEINAMLP STTAISHASF QEKEAGDGST QHIAVSTESV VDDEHLQLDD EETANIDTDT RVIEAEPTWI SFSRPSRIEK SRRARKSSQT ILEKEEATTS ENSNLVSTDE PHLGGVLAAA DVGGKQDATN YNSIFANRFA AFDELLSILK TKFACRVLFE ETLVLPKVGR SRLHLCKDGS PRVIKAVGVQ RNGSEFVLLE VDASDGVKML STKVLSGVDS ETWRNDFEKI RRGVVKSSLN WPNSLFDQLY GQDGHRGVNH PKGLGELQVS REDMEGWAER VVREQFTH |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | ybgA | YbgA | Tn7 | 125 | 8092-8469 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Protein Sequence: | MSAAKALAYA LCDKLESKKV DEVVRNLVSS GWNIREVAWE ASSFDHEMPY FDTGRDFDIG FPDTLSSKIE FLTANGSIVQ ARARLHFKQA LIFAKASKHV AALKNSLDYY YGSSVIPPQN HGGYL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | ybfB | YbfB | Tn7 | 107 | 8523-8846 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Protein Sequence: | MGIAYKLSCH RCGYESDLLY LGQGMAMLPE QIVGTCTCCD NLTTISINES YGVCSLCGAK DRVIFTSYQH EARPRNPCYL ERSFKYQCPR CHAFSMEPPD LPEMLWD |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | ybfC | YbfC | Tn7 | 123 | 8879-9250 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Protein Sequence: | MSYEPPRDGY TNGYIVDVMY DNRPSKYPAV FRTYVRIPDY VELDNAEELA DAIIREVELK EAWLTRVLGN SADPDASVGK WDRRLAQCGG IDEMSSGVQA TYFSPNGQLV GLVAYIDAEW MDE |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | orfX-R721_21 | OrfX-R721_21 | In_Tn7 | 165 | 9311-9808 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Protein Sequence: | MRDLLKVVLS LVLAMASGQA LADLPIVLVD EARLPYDYSP SNYDISPSNY DNSISNYDNS PSNYDNSESN YDNSSSNYDN SRNGNRRLIY SANGSRTFAG YYVIANNGTT NFFSTSGKRM FYTPKGGRGV YGGKDGSFCG ALVVINGQFS LALTDNGLKI MYLSN |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | ANT(3'')-IIa (ARO:3004089) | ANT(3'')-IIa | In_Tn7 | 262 | 9884-10672 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'')(ARO:3004275) | ||||||||||
| Transposase Chemistry: | aminoglycoside nucleotidyltransferase | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3004089 (bitscore: 569)||Synonym: ANT(9)(3'') | ||||||||||
| Protein Sequence: | MREAVIAEVS TQLSEVVGVI ERHLEPTLLA VHLYGSAVDG GLKPHSDIDL LVTVTVRLDE TTRRALINDL LETSASPGES EILRAVEVTI VVHDDIIPWR YPAKRELQFG EWQRNDILAG IFEPATIDID LAILLTKARE HSVALVGPAA EELFDPVPEQ DLFEALNETL TLWNSPPDWA GDERNVVLTL SRIWYSAVTG KIAPKDVAAD WAMERLPAQY QPVILEARQA YLGQEDRLAS RADQLEEFVH YVKGEITKVV GK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | SAT-2 (ARO:3002895) | SAT-2 | In_Tn7 | 174 | 10730-11254 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | streptothricin acetyltransferase (SAT) (ARO:3000869) | ||||||||||
| Target: | nucleoside antibiotic (ARO:3000034) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002895 (bitscore: 365) | ||||||||||
| Protein Sequence: | MKISVIPEQV AETLDAENHF IVREVFDVHL SDQGFELSTR SVSPYRKDYI SDDDSDEDSA CYGAFIDQEL VGKIELNSTW NDLASIEHIV VSHTHRGKGV AHSLIEFAKK WALSRQLLGI RLETQTNNVP ACNLYAKCGF TLGGIDLFTY KTRPQVSNET AMYWYWFSGA QDDA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | dfrA1 (ARO:3002854) | DfrA1 | In_Tn7 | 157 | 11349-11822 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | trimethoprim resistant dihydrofolate reductase dfr (ARO:3001218) | ||||||||||
| Target: | diaminopyrimidine antibiotic (ARO:3000171) | ||||||||||
| Comment: | Synonym: dfr1 | ||||||||||
| Protein Sequence: | MKLSLMVAIS KNGVIGNGPD IPWSAKGEQL LFKAITYNQW LLVGRKTFES MGALPNRKYA VVTRSSFTSD NENVLIFPSI KDALTNLKKI TDHVIVSGGG EIYKSLIDQV DTLHISTIDI EPEGDVYFPE IPSNFRPVFT QDFASNINYS YQIWQKG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI2 | IntI2 | In_Tn7 | 325 | 12154-13131 | + |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 2 | ||||||||||
| Sequence Family: | Class 2 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Comment: | truncated stop at codon 174 | ||||||||||
| Protein Sequence: | MSNSPFLNSI RTDMRQKGYA LKTEKTYLHW IKRFILFHKK RHPQTMGSEE VRLFLSSLAN SRHVAINTQK IALNALAFLY NRFLQQPLGD IDYIPASKPR RLPSVISANE VQRILQVMDT RNQVIFTLLY GAGLRINECL RLRVKDFDFD NGCITVHDGK GGKSRNSLLP TRLIPAIK*L IEQARLIQQD DNLQGVGPSL PFALDHKYPS AYRQAAWMFV FPSSTLCNHP YNGKLCRHHL HDSVARKALK AAVQKAGIVS KRVTCHTFRH SFATHLLQAG RDIRTVQELL GHNDVKTTQI YTHVLGQHFA GTTSPADGLM LLINQ |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| In_Tn7-NC_002525 | In_Tn7 | integron | 9311-13131 | + | 3821 |
1. Datta N, Hedges RW.
Trimethoprim resistance conferred by W plasmids in Enterobacteriaceae.. J Gen Microbiol. 1972 Sep;72(2):349-55. doi: 10.1099/00221287-72-2-349. PubMed ID:
4562309
.
2. Kim SR, Komano T. Nucleotide sequence of the R721 shufflon.. J Bacteriol. 1992 Nov;174(21):7053-8. doi: 10.1128/jb.174.21.7053-7058.1992. PubMed ID: 1400257 .
3. Fennewald MA, Shapiro JA. Transposition of Tn7 in Pseudomonas aeruginosa and isolation of alk::Tn7 mutations.. J Bacteriol. 1979 Jul;139(1):264-9. doi: 10.1128/jb.139.1.264-269.1979. PubMed ID: 110782 .
4. Gringauz E, Orle KA, Waddell CS, Craig NL. Recognition of Escherichia coli attTn7 by transposon Tn7: lack of specific sequence requirements at the point of Tn7 insertion.. J Bacteriol. 1988 Jun;170(6):2832-40. doi: 10.1128/jb.170.6.2832-2840.1988. PubMed ID: 2836374 .
5. Stellwagen AE, Craig NL. Avoiding self: two Tn7-encoded proteins mediate target immunity in Tn7 transposition.. EMBO J. 1997 Nov 17;16(22):6823-34. doi: 10.1093/emboj/16.22.6823. PubMed ID: 9362496 .
6. Waddell CS, Craig NL. Tn7 transposition: two transposition pathways directed by five Tn7-encoded genes.. Genes Dev. 1988 Feb;2(2):137-49. doi: 10.1101/gad.2.2.137. PubMed ID: 2834269 .
7. Skelding Z, Queen-Baker J, Craig NL. Alternative interactions between the Tn7 transposase and the Tn7 target DNA binding protein regulate target immunity and transposition.. EMBO J. 2003 Nov 3;22(21):5904-17. doi: 10.1093/emboj/cdg551. PubMed ID: 14592987 .
8. McKown RL, Waddell CS, Arciszewska LK, Craig NL. Identification of a transposon Tn7-dependent DNA-binding activity that recognizes the ends of Tn7.. Proc Natl Acad Sci U S A. 1987 Nov;84(22):7807-11. doi: 10.1073/pnas.84.22.7807. PubMed ID: 2825163 .
9. Bainton RJ, Kubo KM, Feng JN, Craig NL. Tn7 transposition: target DNA recognition is mediated by multiple Tn7-encoded proteins in a purified in vitro system.. Cell. 1993 Mar 26;72(6):931-43. doi: 10.1016/0092-8674(93)90581-a. PubMed ID: 8384534 .
10. McKown RL, Orle KA, Chen T, Craig NL. Sequence requirements of Escherichia coli attTn7, a specific site of transposon Tn7 insertion.. J Bacteriol. 1988 Jan;170(1):352-8. doi: 10.1128/jb.170.1.352-358.1988. PubMed ID: 2826397 .
11. Arciszewska LK, Drake D, Craig NL. Transposon Tn7. cis-Acting sequences in transposition and transposition immunity.. J Mol Biol. 1989 May 5;207(1):35-52. doi: 10.1016/0022-2836(89)90439-7. PubMed ID: 2544738 .
12. Hansson K, Sundstrom L, Pelletier A, Roy PH. IntI2 integron integrase in Tn7.. J Bacteriol. 2002 Mar;184(6):1712-21. doi: 10.1128/JB.184.6.1712-1721.2002. PubMed ID: 11872723 .
13. Choi KY, Spencer JM, Craig NL. The Tn7 transposition regulator TnsC interacts with the transposase subunit TnsB and target selector TnsD.. Proc Natl Acad Sci U S A. 2014 Jul 15;111(28):E2858-65. doi: 10.1073/pnas.1409869111. Epub 2014 Jun 30. PubMed ID: 24982178 .
14. Wolkow CA, DeBoy RT, Craig NL. Conjugating plasmids are preferred targets for Tn7.. Genes Dev. 1996 Sep 1;10(17):2145-57. doi: 10.1101/gad.10.17.2145. PubMed ID: 8804309 .
15. Rao JE, Craig NL. Selective recognition of pyrimidine motif triplexes by a protein encoded by the bacterial transposon Tn7.. J Mol Biol. 2001 Apr 13;307(5):1161-70. doi: 10.1006/jmbi.2001.4553. PubMed ID: 11292332 .
16. Craig NL. Tn7: a target site-specific transposon.. Mol Microbiol. 1991 Nov;5(11):2569-73. doi: 10.1111/j.1365-2958.1991.tb01964.x. PubMed ID: 1664019 .
17. Chakrabarti A, Desai P, Wickstrom E. Transposon Tn7 protein TnsD binding to Escherichia coli attTn7 DNA and its eukaryotic orthologs.. Biochemistry. 2004 Mar 16;43(10):2941-6. doi: 10.1021/bi035535u. PubMed ID: 15005630 .
18. DeBoy RT, Craig NL. Tn7 transposition as a probe of cis interactions between widely separated (190 kilobases apart) DNA sites in the Escherichia coli chromosome.. J Bacteriol. 1996 Nov;178(21):6184-91. doi: 10.1128/jb.178.21.6184-6191.1996. PubMed ID: 8892817 .
19. Sarnovsky RJ, May EW, Craig NL. The Tn7 transposase is a heteromeric complex in which DNA breakage and joining activities are distributed between different gene products.. EMBO J. 1996 Nov 15;15(22):6348-61. PubMed ID: 8947057 .
20. Peters JE. Tn7.. Microbiol Spectr. 2014 Oct;2(5). doi: 10.1128/microbiolspec.MDNA3-0010-2014. PubMed ID: 26104363 .
21. Kuduvalli PN, Rao JE, Craig NL. Target DNA structure plays a critical role in Tn7 transposition.. EMBO J. 2001 Feb 15;20(4):924-32. doi: 10.1093/emboj/20.4.924. PubMed ID: 11179236 .
22. DeBoy RT, Craig NL. Target site selection by Tn7: attTn7 transcription and target activity.. J Bacteriol. 2000 Jun;182(11):3310-3. doi: 10.1128/JB.182.11.3310-3313.2000. PubMed ID: 10809719 .
23. Sharpe PL, Craig NL. Host proteins can stimulate Tn7 transposition: a novel role for the ribosomal protein L29 and the acyl carrier protein.. EMBO J. 1998 Oct 1;17(19):5822-31. doi: 10.1093/emboj/17.19.5822. PubMed ID: 9755182 .
24. Shi Q, Parks AR, Potter BD, Safir IJ, Luo Y, Forster BM, Peters JE. DNA damage differentially activates regional chromosomal loci for Tn7 transposition in Escherichia coli.. Genetics. 2008 Jul;179(3):1237-50. doi: 10.1534/genetics.108.088161. Epub 2008 Jun 18. PubMed ID: 18562643 .
25. Parks AR, Peters JE. Tn7 elements: engendering diversity from chromosomes to episomes.. Plasmid. 2009 Jan;61(1):1-14. doi: 10.1016/j.plasmid.2008.09.008. Epub 2008 Nov 1. PubMed ID: 18951916 .
26. Gamas P, Craig NL. Purification and characterization of TnsC, a Tn7 transposition protein that binds ATP and DNA.. Nucleic Acids Res. 1992 May 25;20(10):2525-32. doi: 10.1093/nar/20.10.2525. PubMed ID: 1317955 .
27. Arciszewska LK, McKown RL, Craig NL. Purification of TnsB, a transposition protein that binds to the ends of Tn7.. J Biol Chem. 1991 Nov 15;266(32):21736-44. PubMed ID: 1657979 .
28. Gay NJ, Tybulewicz VL, Walker JE. Insertion of transposon Tn7 into the Escherichia coli glmS transcriptional terminator.. Biochem J. 1986 Feb 15;234(1):111-7. doi: 10.1042/bj2340111. PubMed ID: 3010949 .
29. Holder JW, Craig NL. Architecture of the Tn7 posttransposition complex: an elaborate nucleoprotein structure.. J Mol Biol. 2010 Aug 13;401(2):167-81. doi: 10.1016/j.jmb.2010.06.003. Epub 2010 Jun 9. PubMed ID: 20538004 .
30. Peters JE, Craig NL. Tn7 transposes proximal to DNA double-strand breaks and into regions where chromosomal DNA replication terminates.. Mol Cell. 2000 Sep;6(3):573-82. doi: 10.1016/s1097-2765(00)00056-3. PubMed ID: 11030337 .
31. Stellwagen AE, Craig NL. Gain-of-function mutations in TnsC, an ATP-dependent transposition protein that activates the bacterial transposon Tn7.. Genetics. 1997 Mar;145(3):573-85. doi: 10.1093/genetics/145.3.573. PubMed ID: 9055068 .
32. Skelding Z, Sarnovsky R, Craig NL. Formation of a nucleoprotein complex containing Tn7 and its target DNA regulates transposition initiation.. EMBO J. 2002 Jul 1;21(13):3494-504. doi: 10.1093/emboj/cdf347. PubMed ID: 12093750 .
33. May EW, Craig NL. Switching from cut-and-paste to replicative Tn7 transposition.. Science. 1996 Apr 19;272(5260):401-4. doi: 10.1126/science.272.5260.401. PubMed ID: 8602527 .
34. Lu F, Craig NL. Isolation and characterization of Tn7 transposase gain-of-function mutants: a model for transposase activation.. EMBO J. 2000 Jul 3;19(13):3446-57. doi: 10.1093/emboj/19.13.3446. PubMed ID: 10880457 .
35. Stellwagen AE, Craig NL. Analysis of gain-of-function mutants of an ATP-dependent regulator of Tn7 transposition.. J Mol Biol. 2001 Jan 19;305(3):633-42. doi: 10.1006/jmbi.2000.4317. PubMed ID: 11152618 .
36. Gary PA, Biery MC, Bainton RJ, Craig NL. Multiple DNA processing reactions underlie Tn7 transposition.. J Mol Biol. 1996 Mar 29;257(2):301-16. doi: 10.1006/jmbi.1996.0164. PubMed ID: 8609625 .
37. Arciszewska LK, Craig NL. Interaction of the Tn7-encoded transposition protein TnsB with the ends of the transposon.. Nucleic Acids Res. 1991 Sep 25;19(18):5021-9. doi: 10.1093/nar/19.18.5021. PubMed ID: 1656385 .
38. Peters JE, Craig NL. Tn7 recognizes transposition target structures associated with DNA replication using the DNA-binding protein TnsE.. Genes Dev. 2001 Mar 15;15(6):737-47. doi: 10.1101/gad.870201. PubMed ID: 11274058 .
39. Ronning DR, Li Y, Perez ZN, Ross PD, Hickman AB, Craig NL, Dyda F. The carboxy-terminal portion of TnsC activates the Tn7 transposase through a specific interaction with TnsA.. EMBO J. 2004 Aug 4;23(15):2972-81. doi: 10.1038/sj.emboj.7600311. Epub 2004 Jul 15. PubMed ID: 15257292 .
40. Kubo KM, Craig NL. Bacterial transposon Tn7 utilizes two different classes of target sites.. J Bacteriol. 1990 May;172(5):2774-8. doi: 10.1128/jb.172.5.2774-2778.1990. PubMed ID: 2158980 .
41. Peters JE, Craig NL. Tn7: smarter than we thought.. Nat Rev Mol Cell Biol. 2001 Nov;2(11):806-14. doi: 10.1038/35099006. PubMed ID: 11715047 .
42. Shi Q, Straus MR, Caron JJ, Wang H, Chung YS, Guarne A, Peters JE. Conformational toggling controls target site choice for the heteromeric transposase element Tn7.. Nucleic Acids Res. 2015 Dec 15;43(22):10734-45. doi: 10.1093/nar/gkv913. Epub 2015 Sep 17. PubMed ID: 26384427 .
2. Kim SR, Komano T. Nucleotide sequence of the R721 shufflon.. J Bacteriol. 1992 Nov;174(21):7053-8. doi: 10.1128/jb.174.21.7053-7058.1992. PubMed ID: 1400257 .
3. Fennewald MA, Shapiro JA. Transposition of Tn7 in Pseudomonas aeruginosa and isolation of alk::Tn7 mutations.. J Bacteriol. 1979 Jul;139(1):264-9. doi: 10.1128/jb.139.1.264-269.1979. PubMed ID: 110782 .
4. Gringauz E, Orle KA, Waddell CS, Craig NL. Recognition of Escherichia coli attTn7 by transposon Tn7: lack of specific sequence requirements at the point of Tn7 insertion.. J Bacteriol. 1988 Jun;170(6):2832-40. doi: 10.1128/jb.170.6.2832-2840.1988. PubMed ID: 2836374 .
5. Stellwagen AE, Craig NL. Avoiding self: two Tn7-encoded proteins mediate target immunity in Tn7 transposition.. EMBO J. 1997 Nov 17;16(22):6823-34. doi: 10.1093/emboj/16.22.6823. PubMed ID: 9362496 .
6. Waddell CS, Craig NL. Tn7 transposition: two transposition pathways directed by five Tn7-encoded genes.. Genes Dev. 1988 Feb;2(2):137-49. doi: 10.1101/gad.2.2.137. PubMed ID: 2834269 .
7. Skelding Z, Queen-Baker J, Craig NL. Alternative interactions between the Tn7 transposase and the Tn7 target DNA binding protein regulate target immunity and transposition.. EMBO J. 2003 Nov 3;22(21):5904-17. doi: 10.1093/emboj/cdg551. PubMed ID: 14592987 .
8. McKown RL, Waddell CS, Arciszewska LK, Craig NL. Identification of a transposon Tn7-dependent DNA-binding activity that recognizes the ends of Tn7.. Proc Natl Acad Sci U S A. 1987 Nov;84(22):7807-11. doi: 10.1073/pnas.84.22.7807. PubMed ID: 2825163 .
9. Bainton RJ, Kubo KM, Feng JN, Craig NL. Tn7 transposition: target DNA recognition is mediated by multiple Tn7-encoded proteins in a purified in vitro system.. Cell. 1993 Mar 26;72(6):931-43. doi: 10.1016/0092-8674(93)90581-a. PubMed ID: 8384534 .
10. McKown RL, Orle KA, Chen T, Craig NL. Sequence requirements of Escherichia coli attTn7, a specific site of transposon Tn7 insertion.. J Bacteriol. 1988 Jan;170(1):352-8. doi: 10.1128/jb.170.1.352-358.1988. PubMed ID: 2826397 .
11. Arciszewska LK, Drake D, Craig NL. Transposon Tn7. cis-Acting sequences in transposition and transposition immunity.. J Mol Biol. 1989 May 5;207(1):35-52. doi: 10.1016/0022-2836(89)90439-7. PubMed ID: 2544738 .
12. Hansson K, Sundstrom L, Pelletier A, Roy PH. IntI2 integron integrase in Tn7.. J Bacteriol. 2002 Mar;184(6):1712-21. doi: 10.1128/JB.184.6.1712-1721.2002. PubMed ID: 11872723 .
13. Choi KY, Spencer JM, Craig NL. The Tn7 transposition regulator TnsC interacts with the transposase subunit TnsB and target selector TnsD.. Proc Natl Acad Sci U S A. 2014 Jul 15;111(28):E2858-65. doi: 10.1073/pnas.1409869111. Epub 2014 Jun 30. PubMed ID: 24982178 .
14. Wolkow CA, DeBoy RT, Craig NL. Conjugating plasmids are preferred targets for Tn7.. Genes Dev. 1996 Sep 1;10(17):2145-57. doi: 10.1101/gad.10.17.2145. PubMed ID: 8804309 .
15. Rao JE, Craig NL. Selective recognition of pyrimidine motif triplexes by a protein encoded by the bacterial transposon Tn7.. J Mol Biol. 2001 Apr 13;307(5):1161-70. doi: 10.1006/jmbi.2001.4553. PubMed ID: 11292332 .
16. Craig NL. Tn7: a target site-specific transposon.. Mol Microbiol. 1991 Nov;5(11):2569-73. doi: 10.1111/j.1365-2958.1991.tb01964.x. PubMed ID: 1664019 .
17. Chakrabarti A, Desai P, Wickstrom E. Transposon Tn7 protein TnsD binding to Escherichia coli attTn7 DNA and its eukaryotic orthologs.. Biochemistry. 2004 Mar 16;43(10):2941-6. doi: 10.1021/bi035535u. PubMed ID: 15005630 .
18. DeBoy RT, Craig NL. Tn7 transposition as a probe of cis interactions between widely separated (190 kilobases apart) DNA sites in the Escherichia coli chromosome.. J Bacteriol. 1996 Nov;178(21):6184-91. doi: 10.1128/jb.178.21.6184-6191.1996. PubMed ID: 8892817 .
19. Sarnovsky RJ, May EW, Craig NL. The Tn7 transposase is a heteromeric complex in which DNA breakage and joining activities are distributed between different gene products.. EMBO J. 1996 Nov 15;15(22):6348-61. PubMed ID: 8947057 .
20. Peters JE. Tn7.. Microbiol Spectr. 2014 Oct;2(5). doi: 10.1128/microbiolspec.MDNA3-0010-2014. PubMed ID: 26104363 .
21. Kuduvalli PN, Rao JE, Craig NL. Target DNA structure plays a critical role in Tn7 transposition.. EMBO J. 2001 Feb 15;20(4):924-32. doi: 10.1093/emboj/20.4.924. PubMed ID: 11179236 .
22. DeBoy RT, Craig NL. Target site selection by Tn7: attTn7 transcription and target activity.. J Bacteriol. 2000 Jun;182(11):3310-3. doi: 10.1128/JB.182.11.3310-3313.2000. PubMed ID: 10809719 .
23. Sharpe PL, Craig NL. Host proteins can stimulate Tn7 transposition: a novel role for the ribosomal protein L29 and the acyl carrier protein.. EMBO J. 1998 Oct 1;17(19):5822-31. doi: 10.1093/emboj/17.19.5822. PubMed ID: 9755182 .
24. Shi Q, Parks AR, Potter BD, Safir IJ, Luo Y, Forster BM, Peters JE. DNA damage differentially activates regional chromosomal loci for Tn7 transposition in Escherichia coli.. Genetics. 2008 Jul;179(3):1237-50. doi: 10.1534/genetics.108.088161. Epub 2008 Jun 18. PubMed ID: 18562643 .
25. Parks AR, Peters JE. Tn7 elements: engendering diversity from chromosomes to episomes.. Plasmid. 2009 Jan;61(1):1-14. doi: 10.1016/j.plasmid.2008.09.008. Epub 2008 Nov 1. PubMed ID: 18951916 .
26. Gamas P, Craig NL. Purification and characterization of TnsC, a Tn7 transposition protein that binds ATP and DNA.. Nucleic Acids Res. 1992 May 25;20(10):2525-32. doi: 10.1093/nar/20.10.2525. PubMed ID: 1317955 .
27. Arciszewska LK, McKown RL, Craig NL. Purification of TnsB, a transposition protein that binds to the ends of Tn7.. J Biol Chem. 1991 Nov 15;266(32):21736-44. PubMed ID: 1657979 .
28. Gay NJ, Tybulewicz VL, Walker JE. Insertion of transposon Tn7 into the Escherichia coli glmS transcriptional terminator.. Biochem J. 1986 Feb 15;234(1):111-7. doi: 10.1042/bj2340111. PubMed ID: 3010949 .
29. Holder JW, Craig NL. Architecture of the Tn7 posttransposition complex: an elaborate nucleoprotein structure.. J Mol Biol. 2010 Aug 13;401(2):167-81. doi: 10.1016/j.jmb.2010.06.003. Epub 2010 Jun 9. PubMed ID: 20538004 .
30. Peters JE, Craig NL. Tn7 transposes proximal to DNA double-strand breaks and into regions where chromosomal DNA replication terminates.. Mol Cell. 2000 Sep;6(3):573-82. doi: 10.1016/s1097-2765(00)00056-3. PubMed ID: 11030337 .
31. Stellwagen AE, Craig NL. Gain-of-function mutations in TnsC, an ATP-dependent transposition protein that activates the bacterial transposon Tn7.. Genetics. 1997 Mar;145(3):573-85. doi: 10.1093/genetics/145.3.573. PubMed ID: 9055068 .
32. Skelding Z, Sarnovsky R, Craig NL. Formation of a nucleoprotein complex containing Tn7 and its target DNA regulates transposition initiation.. EMBO J. 2002 Jul 1;21(13):3494-504. doi: 10.1093/emboj/cdf347. PubMed ID: 12093750 .
33. May EW, Craig NL. Switching from cut-and-paste to replicative Tn7 transposition.. Science. 1996 Apr 19;272(5260):401-4. doi: 10.1126/science.272.5260.401. PubMed ID: 8602527 .
34. Lu F, Craig NL. Isolation and characterization of Tn7 transposase gain-of-function mutants: a model for transposase activation.. EMBO J. 2000 Jul 3;19(13):3446-57. doi: 10.1093/emboj/19.13.3446. PubMed ID: 10880457 .
35. Stellwagen AE, Craig NL. Analysis of gain-of-function mutants of an ATP-dependent regulator of Tn7 transposition.. J Mol Biol. 2001 Jan 19;305(3):633-42. doi: 10.1006/jmbi.2000.4317. PubMed ID: 11152618 .
36. Gary PA, Biery MC, Bainton RJ, Craig NL. Multiple DNA processing reactions underlie Tn7 transposition.. J Mol Biol. 1996 Mar 29;257(2):301-16. doi: 10.1006/jmbi.1996.0164. PubMed ID: 8609625 .
37. Arciszewska LK, Craig NL. Interaction of the Tn7-encoded transposition protein TnsB with the ends of the transposon.. Nucleic Acids Res. 1991 Sep 25;19(18):5021-9. doi: 10.1093/nar/19.18.5021. PubMed ID: 1656385 .
38. Peters JE, Craig NL. Tn7 recognizes transposition target structures associated with DNA replication using the DNA-binding protein TnsE.. Genes Dev. 2001 Mar 15;15(6):737-47. doi: 10.1101/gad.870201. PubMed ID: 11274058 .
39. Ronning DR, Li Y, Perez ZN, Ross PD, Hickman AB, Craig NL, Dyda F. The carboxy-terminal portion of TnsC activates the Tn7 transposase through a specific interaction with TnsA.. EMBO J. 2004 Aug 4;23(15):2972-81. doi: 10.1038/sj.emboj.7600311. Epub 2004 Jul 15. PubMed ID: 15257292 .
40. Kubo KM, Craig NL. Bacterial transposon Tn7 utilizes two different classes of target sites.. J Bacteriol. 1990 May;172(5):2774-8. doi: 10.1128/jb.172.5.2774-2778.1990. PubMed ID: 2158980 .
41. Peters JE, Craig NL. Tn7: smarter than we thought.. Nat Rev Mol Cell Biol. 2001 Nov;2(11):806-14. doi: 10.1038/35099006. PubMed ID: 11715047 .
42. Shi Q, Straus MR, Caron JJ, Wang H, Chung YS, Guarne A, Peters JE. Conformational toggling controls target site choice for the heteromeric transposase element Tn7.. Nucleic Acids Res. 2015 Dec 15;43(22):10734-45. doi: 10.1093/nar/gkv913. Epub 2015 Sep 17. PubMed ID: 26384427 .