Mobile element type:
Transposon
Name:
Tn558.3
Synonyms:
Accession:
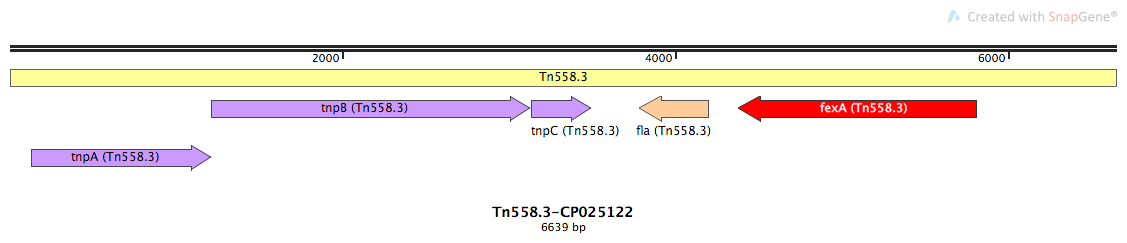
Tn558.3-CP025122
Family:
Tn554
Group:
First isolate:
Yes
Partial:
ND
Evidence of transposition:
Yes
Host Organism:
Bacillus sp. HBCD-sjtu
Date of Isolation:
2017
Country:
China
Molecular Source:
chromosome
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tnpA | Tn558.3 | 132-1217 | + | Transposase | |
| tnpB | Tn558.3 | 1214-3133 | + | Transposase | |
| tnpC | Tn558.3 | 3135-3500 | + | Accessory Gene | Helper |
| fla | Tn558.3 | 3783-4199 | - | Passenger Gene | Other |
| fexA (ARO:3002704) | Tn558.3 | 4377-5804 | - | Passenger Gene | Antibiotic Resistance |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn558.3 | 361 | 132-1217 | + |
|---|
| Class: | Transposase | ||||||||||
| Comment: | tyrosine recombinase XerD||TIGR02225||phage integrase, N-terminal SAM-like domain||DNA breaking-rejoining enzymes C-terminal catalytic domain | ||||||||||
| Protein Sequence: | MKVQKVIVEE KSYPLYILLD KEFEVIEPVK RFIKYLDNTG KAPNTIKTYC YHLKLFYEFM SQSRIELGDL QFEDLANFVG WLRNPAGHLK VIDIQPKKAK REETSVNSIL NAVTSFLEYL NRTENFKAID MSKEARGRNF KGFLHHISKG RSYKKNILKL RVKKKLVQVL EHGQVKAIIE ACHTKRDKLL IMLMYEGGLR IGEALSLRIE DISTWDNQIN IRPRDHNENG AYIKLKKERT IDVSKELMAL YTDYLVHEYG EDLDHDYIFI NLKDSYFGHP LKYQSVLDLI RRLGKRTGIT FTAHILRHTH ATELIRSGWD AAYVQKRLGH AHVQTTLDTY VHLSDQDMKN EYKKYLQGRE T |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpB | TnpB | Tn558.3 | 639 | 1214-3133 | + |
|---|
| Class: | Transposase | ||||||||||
| Comment: | Phage integrase N-terminal SAM-like domain||Region: Phage_int_SAM_1||cl12235||Phage integrase family||pfam00589 | ||||||||||
| Protein Sequence: | MTMKFPSVAT NRKLIASTEI NKKIANMTSV LKGYWAADRW DIRICPHPSA IELSKNPSLR NRWVNFDKVE NVWLKTELKY FYYLHLNNGA WNAKSVWIRK GTVISRMMGF LNLKYPNITS ITEVPIKKAL TEYRTYLTEQ KVKTTTTNYK LDVNQQKVTV HANSYYVTHL KQFMEFYEDF YFDGEEWEKD VWNRRKLSLP EDKVNPTSYE YTINFKGFKN NYFKEIVKRY CKLMLNTASF SHVVDIASKL KEFFNFMNKN CEGIQRIHQL TRNEIEQYFN YINLKGLKPS TVTGRISTLD VFFTTIQRYD WKDTPSKILI FQEDYPKVPK ALPRYIDEHI LEQLNGKLDK LEPYIATMVM VLQECGMRIS ELCTLKKGSV ITDKEGDCFL KYYQWKMKKE HIIPISKEIA ALILVQEQRV ADELDDGCVY VFPRKDCSPL KQDTFRVKLN ELAYEEKITD SNGEIFRFHA HAFRHTVGTR MINNGVPQHI VQKFLGHESP EMTARYAHIF DETLKKEFTK FKETLVTNNG SILDLSEENT EADNTDLQWF KKNINAQALP NGYCRLPVIA GPCPHANACL DCTNFCTSKQ FLTEHEEHLE RTKEILNRAK QNQWQRQVET NERVKNRLEQ IIHSLKETN |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpC | TnpC | Tn558.3 | 121 | 3135-3500 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Helper | ||||||||||
| Sequence Family: | Tn554_family | ||||||||||
| Comment: | target specificity and orientation in Tn554 family | ||||||||||
| Protein Sequence: | MIRKGNTTAI VQLAKDKSEK TRIRVEKTIS EMALKEEKIN FNSVAQKANV SKSWLYKQKD IRTRVETLRG MQISELTPRK PSKSPRSEDV LIKTLKSRIK ALEEENERLK DQVQKLHGKL F |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | fla | Fla | Tn558.3 | 138 | 3783-4199 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | flavodoxin | ||||||||||
| Protein Sequence: | MKTLVIVSHP DMPNSRINKV WVEKAASYSN EITIHDLYRE YPDFIINVKR EQELVENHDN IIFQFPLYWY SSPSLLKKWI DEVIIYGWAY GSKGKRIFYN RKLGLAISAG VKKGEFTSMG KNKHTLTQML TPFKSLCA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | fexA (ARO:3002704) | FexA | Tn558.3 | 475 | 4377-5804 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | phenicol antibiotic (ARO:3000387) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002704 | ||||||||||
| Protein Sequence: | MKKDSKSKEM IQSEKRGSTR LLMMVLSLSV LVAAITVDLV NPVLPLISKD LEASKSQVSW IVSGIALVLA IGVPIYGRIS DFFELRKLYI FAIMILASGS LLCAIAPNLP LLVLGRMVQG AGMSAIPVLS VIAISKVFPQ GKRGGALGII AGSIGVGTAA GPIFGGVVGQ YLGWNALFWF TFLLAIMIVI GAYYALPTIK PAESVGSNKN FDFIGGLFLG LTVGLLLFGI TQGETSGFSS FSSLTSLIGS VVALVGFIWR IVTAENPFVP PVLFNNKDYV NTVIIAFFSM FAYFAVLVFV PLLVVEVNGL SSGQAGMILL PGGVAVAILS PFVGRLSDRF GDKRLIITGM TLMGLSTLFL STYASGASPL LVSVGVLGVG IAFAFTNSPA NNAAVSALDA DKVGVGMGIF QGALYLGAGT GAGMIGALLS ARRDATEPIN PLYILDAMSY SDAFLAATGA ILIALIAGLG LKKRG |
||||||||||