Mobile element type:
Transposon
Name:
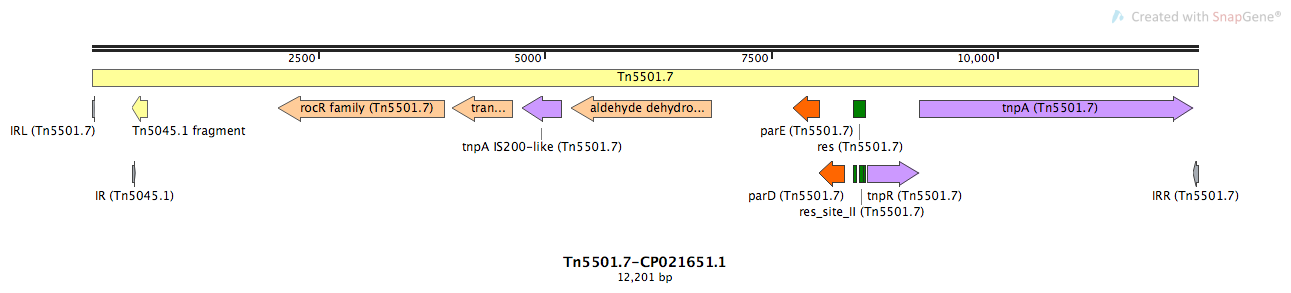
Tn5501.7
Synonyms:
Accession:
Tn5501.7-CP021651.1
Family:
Tn3
Group:
Tn3000
First isolate:
Yes
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Acidovorax sp. T1
Date of Isolation:
2017
Country:
China
Molecular Source:
plasmid p3-T1
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGTTCTAA GCCGGAACCG CCGAAAATTC CGTCAGCC |
| IRR | 12164-12201 | - | 38 | GGGGTTCTAA GCCAGAACCG CCGAAATTTC CGTCATCC |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| res | Tn5501.7 | 8412-8542 | + | 131 | GCCTGTCGGA AAACATTTGT TTTTCGACAG GCCTTCAACG GTCCTCTGCA CCAACCTCCG AGTGGCCGCA AAATTGTGCG GAAAACTCTG TCGCCAGACG CTACCATACG GAAACCTCGT CTTAATGGTT T |
| res_site_I | Tn5501.7 | 8412-8440 | + | 29 | GCCTGTCGGA AAACATTTGT TTTTCGACA |
| res_site_II | Tn5501.7 | 8474-8517 | + | 44 | TGGCCGCAAA ATTGTGCGGA AAACTCTGTC GCCAGACGCT ACCA |
| res_site_III | Tn5501.7 | 8518-8542 | + | 25 | TACGGAAACC TCGTCTTAAT GGTTT |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| rocR family | Tn5501.7 | 2060-3889 | - | Passenger Gene | Other |
| transporter | Tn5501.7 | 3986-4639 | - | Passenger Gene | Other |
| tnpA IS200-like | Tn5501.7 | 4757-5188 | - | Transposase | |
| aldehyde dehydrogenase | Tn5501.7 | 5293-6840 | - | Passenger Gene | Other |
| parE | Tn5501.7 | 7752-8036 | - | Passenger Gene | Toxin |
| parD | Tn5501.7 | 8033-8311 | - | Passenger Gene | Antitoxin |
| tnpR | Tn5501.7 | 8565-9143 | + | Accessory Gene | Resolvase |
| tnpA | Tn5501.7 | 9140-12169 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | rocR family | RocR family | Tn5501.7 | 609 | 2060-3889 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Comment: | sigma-54-dependent Fis family transcriptional regulator | ||||||||||
| Protein Sequence: | MDAHTLHDQQ RLVIARQRFV EGVALPSNLL PSTIAHSWAR SREAGLLPWQ SWLAEQDDSL LPLDESDQQL AECVRPEIDR LWQQIGGASW TIFCVNTRGV IVHARQARTL DGPLLPLQIG RRIQESDIGT TAPSCTLAEG VPMVLIGNQH YLSDFEHFFC VSVPIRGLHG EVIGVLDITG IGDRQAGAVL EQLSHAAMAA ENRFFTTLGD CRILSLQHDP RLLGTPLQGL LAVDQTGRIQ NANRAAQRLL GLEQFRPIGK RLDQLFEESS AIDLHRAQQQ LTLSDGSRLY AQVIEQRRES TALHVCSPLL CVSILGADPQ LNRQLDNARK AFAAGVPILL QGETGTGKEV FARALHDQWN AQAPFVAINC SAIPESLIEA ELFGYVEGTF TGARKGGASG QLEAAHGGTL LLDEIGDMPA ALQTRLLRVL QEREITRLGS TTRRALDVRV ISATHCDLQQ RMLDKSFRED LYYRLNGLKI RLPALRERQD LNTLIDLLRQ RYGAPPLEHQ ALMALHRQVW PGNIRQLEQT LRLAAALASD QPMILLEHLP ADMRAESEQE RGSDLQDAVR RTVESALQAN AGNISATASQ LGISRTTLYK KLGRITKSE |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | transporter | Transporter | Tn5501.7 | 217 | 3986-4639 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Protein Sequence: | MDIGGMTVDP QILLPFGHVY DARIAGQSLG SASGLADPII GATFWLVNQP AAGASGRYFG ITPLLYLPWG SYDKHDTLNL GENRFKGDLQ LGWVEPLWGK VSMELYGDAV VYGHNDDAGT GNQTLKQDAT YQLQGNLRYD FNPAQRVALG YSTSTGGKQY LDGNYTGQKT EVQQVRAEFQ QMVGRTVQLS AQLTHDIHVV GGFQEDIGVN LRALLLF |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA IS200-like | TnpA IS200-like | Tn5501.7 | 143 | 4757-5188 | - |
|---|
| Class: | Transposase | ||||||||||
| Function: | single-strand transposase | ||||||||||
| Transposase Chemistry: | HUH | ||||||||||
| Protein Sequence: | MQDYQSLSHT KWDCKYHVVF IPKKRKKLIF GAIRKHLGEV LHELAGQKEC RILEGHLMSD HIHMCISIPP KYSVANVVGY IKGKSAILIA RRFNGRSRNF GGENFWARGY FVSTVGLDEE MVRAYIRHQE KEDEHYDQLK LGM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aldehyde dehydrogenase | Aldehyde dehydrogenase | Tn5501.7 | 515 | 5293-6840 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Protein Sequence: | MFFIAANDGW PPNHKIKKRG IYPMSSTSLP AIFLPAELWN DRLFTGTWEA GPIAATQVLE PATGNLLGYV ANADAARVAA SAKTASAAQT SWHQQPYDER AALFRKAAKI AEQHFEAIVD WLVRESGSTR AKAAFETSIS IKVLHESAGL PSRSQGEVLP SVPGRLSLAR RRPLGVVGVI SPFNFPLYLA IRAVAPALAT GNAVVLKPDP RTAVCGGFVI ARLFELAGLP AGLLHVLPGG VDAGAALTSD PHVAMIQFTG STGAGRKVGE AAGRHLKKVS LELGGKNSLI VLDDADLDLA LANATWGVYL HQGQICMSTG RLLVQRAIYD RFLERLVAKA KSLTVGDPAT SEVHLGPLIN AQQRDHAHRV VQAAQQAGAT LEAGGGYEGL FFEPTVLGGV TRDNPAFNEE IFGPVAVVVP FDTDEEAVAL ANDTAYGLSA AVLSRDVGRA LKLGEQLRVG LLHINDQTVN DEVINPFGGV GASGNGTSIG GAANWEEFTQ WQWLTVKGEA PAYPL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | parE | ParE | Tn5501.7 | 94 | 7752-8036 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Toxin | ||||||||||
| Sequence Family: | ParE_toxin (Pfam:PF05016) | ||||||||||
| Target: | DNA gyrase | ||||||||||
| Protein Sequence: | VRVVWTPEAQ QDRADVWDYI AADNPRAAAR MDEIFSDAAA RLIQHPMLGK PGKIPGTREL IPHESYRLVY QIDGETVWIL TLVHTARLWP PVRD |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | parD | ParD | Tn5501.7 | 92 | 8033-8311 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antitoxin | ||||||||||
| Sequence Family: | parD (PDB:4Q2U) | ||||||||||
| Comment: | RelB/ParD/CcdA/DinJ | ||||||||||
| Protein Sequence: | MSKQAVFTMK LEPELRAEFM AEAEAAHRPA SQVLRELMRE FVQRQRESRE YDEFLRRKVE AGRASMRAGL GRSNDEVEAE FAARRASVAS QA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | Tn5501.7 | 192 | 8565-9143 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MRVSSDSDRQ STNLQRDALL AVGVDARHLF EDHASGAKDD RAGLARALEF VRPGDVLVVW KLDRLGRSLS HLLAIVTSLK KKQVAFRSLT ENLDTTTPSG EFLFQVFGAL AQYERALIQE RVVAGLAAAR KRGRIGGRPQ AITGEKLEAI VAALDGGMSK AAVCRNFGVK RTTLIETLAR VGWTGSRGAS SR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn5501.7 | 1009 | 9140-12169 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MTTKSERLTV LSDAEQEALY GLPDFDDAQR LEYLALTETE LALASSRPGL HAQVYCILQI GYFKAKHAFF RFDWSEVEHD CAFVLSRYFH GESFEHKPIS KHEHYTQREW IADLFGYRPW AAEFLAQLAQ QAAQTVRRDV MPGFIAAELI VWLNEHKIIR PGYTTLQELV SEALSAERRR LAGLLSEVLD ESAKAALGRL LVRDDTLSQL AALKQDAKDF GWRQMARERE KRATLEPLHR IAKALLPKLG VSQQNLLYYA SLANFYTVHD LRNLKADQTY LYLLCYAWVR YRQLSDNLVD AMAYHMKQLE DESSAGAKQS FVAEQVRRQQ DTPQVGRLLS LYIDDSVPDP TPFGDVRQRA YKIMPRDTLQ TTAQRMSVKP VSKLALHWQA VDGLAERIRR HLRPLYVALD LAGTDPGSPW LVALAWAKDV FAKQQRLSQR PLAECPAATL PKRLRPYLLT FDADGKPTDL HADRYEFWLY RQVRKRFQSG ELYLDDSLQH RHFSDELVSL DEKAAVLAQI DIPFLRQPLD AQLDALATEL RAQWLAFNRE LKQGKLTHLE YDKDTQKLTW RKPKGENQKA REKAFYEQLP FCDVADVFRF VNGQCQFLSA LTPLQPRYAK KVADADSLMA VIIAQAMNHG NQVMARTSDI PYHVLESAYQ QYLRHATLHA ANDCISNAIA ALPIFPYYSF DLDALYGAVD GQKFGVERPT VKARHSRKYF GRGKGVVAYT LLCNHVPLNG YLIGAHDYEA HHVFDIWYRN TSDIVPTAIT GDMHSVNKAN FAILHWFGLR FEPRFTDLGD QLKELYSADD PALYDQCLIR PAGRIDRDLI VSEKPNLDQI VATLGLKEMT QGTLIRKLCT YTAPNPTRRA VFEFDKLIRS IYTLRYLRDP QLERNVHRSQ NRIESYHQLR STIAQVGGKK ELTGRTDIEI EISNQCARLI ANAVIFYNSA ILSRLLMKYE ASGNAKAHAL LTQISPAAWR HILLNGHYTF QSDGKMIDLD ALVAGLELG |
||||||||||
1. Chu CW, Liu B, Li N, Yao SG, Cheng D, Zhao JD, Qiu JG, Yan X, He Q, He J.
A Novel Aerobic Degradation Pathway for Thiobencarb Is Initiated by the TmoAB Two-Component Flavin Mononucleotide-Dependent Monooxygenase System in Acidovorax sp. Strain T1.. Appl Environ Microbiol. 2017 Nov 16;83(23):e01490-17. doi: 10.1128/AEM.01490-17. Print 2017 Dec 1. PubMed ID:
28939603
.