Mobile element type:
Transposon
Name:
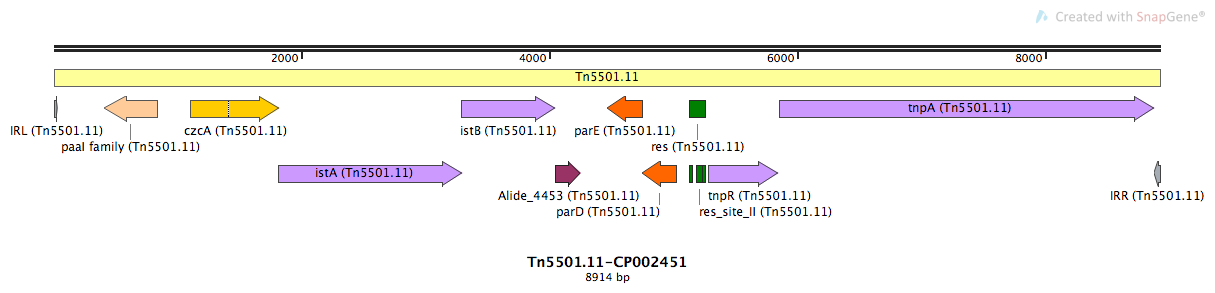
Tn5501.11
Synonyms:
Accession:
Tn5501.11-CP002451
Family:
Tn3
Group:
Tn3000
First isolate:
Yes
Partial:
ND
Evidence of transposition:
Yes
Host Organism:
Alicycliphilusdenitrificans BC
Date of Isolation:
2011
Country:
Korea
Molecular Source:
plasmid pALIDE02
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGTTCTAA GCCGGAACCG CCGAAAATTC CGTCAGCC |
| IRR | 8877-8914 | - | 38 | GGGGTTCTAA GCCGGAACCG CCGAAATTTC CGTCATCC |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| res | Tn5501.11 | 5125-5255 | + | 131 | GCCTGTCGGA AAACATTTGT TTTTCGACAG GCCTTCAACG ATCCTCTGCA CCAACCTCCG AGTGGCCGCA AAATTGTGCG GAAAACTCTG TCGCCAGACG CTACCATACG GAAACCTCGT CTTAATGGTT T |
| res_site_I | Tn5501.11 | 5125-5153 | + | 29 | GCCTGTCGGA AAACATTTGT TTTTCGACA |
| res_site_II | Tn5501.11 | 5187-5230 | + | 44 | TGGCCGCAAA ATTGTGCGGA AAACTCTGTC GCCAGACGCT ACCA |
| res_site_III | Tn5501.11 | 5231-5255 | + | 25 | TACGGAAACC TCGTCTTAAT GGTTT |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| paaI family | Tn5501.11 | 405-833 | - | Passenger Gene | Other |
| czcA | Tn5501.11 | 1104-1831 | + | Passenger Gene | Heavy Metal Resistance |
| czcA N-ter | Tn5501.11 | 1104-1400 | + | Passenger Gene | Heavy Metal Resistance |
| czcA C-ter | Tn5501.11 | 1400-1831 | + | Passenger Gene | Heavy Metal Resistance |
| istA | Tn5501.11 | 1815-3305 | + | Transposase | |
| istB | Tn5501.11 | 3292-4050 | + | Accessory Gene | ATPase Transposition Helper |
| Alide_4453 | Tn5501.11 | 4047-4259 | + | Passenger Gene | Hypothetical |
| parE | Tn5501.11 | 4465-4749 | - | Passenger Gene | Toxin |
| parD | Tn5501.11 | 4746-5024 | - | Passenger Gene | Antitoxin |
| tnpR | Tn5501.11 | 5278-5856 | + | Accessory Gene | Resolvase |
| tnpA | Tn5501.11 | 5853-8882 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | paaI family | PaaI family | Tn5501.11 | 142 | 405-833 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Comment: | PaaI family thioesterase | ||||||||||
| Protein Sequence: | MSKLRSDVSL ETLNERGKEY LPGYLGIQVI DLQPNVLRSR MPIQKLHIAP NEFLHAASVV ALADTSCGYA TIAHLPDGAQ SFTTIELKSN HLGTVTEGTV ACVATAQHLG RTTQVWDAEV TDEASGRKLA LFRCTQMILW PR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | czcA | CzcA | Tn5501.11 | 242 | 1104-1831 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Comment: | heavy metal efflux pump, CzcA family || split by frameshift mutation | ||||||||||
| Protein Sequence: | MGSGALEGAR RATGNAPDPG APMAGGVFVQ SGGRAFGTRV ADARTIGLGL WQRRRDTGSN RQVIPSVAPH TRARLRGRCA RSLSRSGRWR RVSHGPQGPW SRPDDAISVP RAARQC*SAA TVIGGSAIAP PAAPSVPARP RCGQPARATR RALRGAWRTR NASAATARDA RK*RIRVPRR RPRLLK*GPR RPCPSPYRPR AGSATCATAH CPRSCVRTSC TPRFGAPVRD PNGVRPAMAL SP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | czcA N-ter | CzcA N-ter | Tn5501.11 | 99 | 1104-1400 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Cobalt||Cadmium||Nickel | ||||||||||
| Comment: | heavy metal efflux pump, CzcA family || split by frameshift mutation||PMID:12829273 | ||||||||||
| Protein Sequence: | MGSGALEGAR RATGNAPDPG APMAGGVFVQ SGGRAFGTRV ADARTIGLGL WQRRRDTGSN RQVIPSVAPH TRARLRGRCA RSLSRSGRWR RVSHGPQGP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | czcA C-ter | CzcA C-ter | Tn5501.11 | 143 | 1400-1831 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Cobalt||Cadmium||Nickel | ||||||||||
| Comment: | CzcA family heavy metal efflux pump || split by frameshift mutation||PMID:12829273 | ||||||||||
| Protein Sequence: | MESTGRRYLC AACRAAVLIC SHCDRGQRYC AAGCAQRARA ASLRAAGTRY QASFAGRLAH AQRQRRYRAR RQEVTHQGSP PPAPPVEVRT APTVPESVPS ASGQCHLCHR ALSPFVRQDF LHSPVRRPCP RSERSPPRHG PVP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | istA | IstA | Tn5501.11 | 496 | 1815-3305 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MALSPEHEAQ ILRYYHAERW RIGTIAVQLG LHRDTVARVL AQAGLPRHGP VQRASAIDPY LPFLHETLAQ FPRLTAARLY DMVRARGYPG RPDHFRHLIA RHRPRPSAEA YLRLRTLPGE QAQVDWGHFG HLTIGRARRP LMAFVMVLSW SRQIYLRFFL DARMENFLRG HVGAFECWGA VPRIALYDNL KSAVLERCGN AIRFHPTLLA LAGHYRFEPR PVAVARGNEK GRVERAIRYV REAFFAGCAF ADLDDLNAQA QAWCEGAAGA RRCPEDASMT VAEAFAAERE RLLARPETPF PTDELRAVSA GKTPYVRFDL NDYSIPHTHV RRTLTVLADP LRVRILDGED VIATHARSYD RRQQIECDAH LEALVAHKHA ASAHRATDRL TAAVPACQAL LAQAAERGEP LGRTTRALTD LLDRYGAGEL AVAVDEALAR GVPHPNAVRL ALERRREAPP PLGVPLPAHL KTRDVTVRAH PLAGYDRLLE DDHDDA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | istB | IstB | Tn5501.11 | 252 | 3292-4050 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | ATPase Transposition Helper | ||||||||||
| Protein Sequence: | MTTPEPLLAR ARALQLHGVV SHWAECAQAP WITPLIEWEE TERARRSLER RLRCAHIGRF KPLADFDWRW PEQCDQAAIA ELMTLGFLET ATNALLVGPS GLGKTTIAQN IAHQAVLRGH TVLFTTAGQL LGELAALDSD SALRRRLRHY ASPTLLVIDE VGYLSYSNRH ADLLFELISR RHEKKSTLIT TNKSFAEWSE VFPNAACVVA LIDRLIHHAE IIALKGPSWR HKEAQERAAT NARKRKSARS PA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | Alide_4453 | Alide_4453 | Tn5501.11 | 70 | 4047-4259 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Protein Sequence: | MKTDHPPSGL LRGLDLLIDA DWTPEQARAV IELLDDLRER IWAHYELALF EHYRQDRLTQ AELPFDDPSF |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | parE | ParE | Tn5501.11 | 94 | 4465-4749 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Toxin | ||||||||||
| Sequence Family: | ParE_toxin (Pfam:PF05016) | ||||||||||
| Target: | DNA gyrase | ||||||||||
| Protein Sequence: | VRVVWTPEAQ QDRADVWDYI AADNSRAAAR MDEIFSDAAA RLIQHPMLGK PGKIPGTREL IPHESYRLVY QIDGETVWIL TLVHTARLWP PVRD |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | parD | ParD | Tn5501.11 | 92 | 4746-5024 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antitoxin | ||||||||||
| Sequence Family: | parD (PDB:4Q2U) | ||||||||||
| Comment: | RelB/ParD/CcdA/DinJ | ||||||||||
| Protein Sequence: | MSKQAVFTMK LEPELRAEFM AEAEAAHRPA SQVLRELMRE FVQRQRESRE YDEFLRRKVE AGRASMRAGL GRSNDEVEAE FAARRASVAS QA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | Tn5501.11 | 192 | 5278-5856 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MRVSSDSDRQ STNLQRDALL AVGVDARHLF EDHASGAKDD RAGLARALEF VRPGDVLVVW KLDRLGRSLS HLLAIVTSLK EKQVAFRSLT ENLDTTTPSG EFLFQVFGAL AQYERALIQE RVVAGLAAAR KRGRIGGRPQ AITGEKLEAI VAALDGGMSK AAVCRNFGVK RTTLIETLAR VGWTGSRGAS SR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn5501.11 | 1009 | 5853-8882 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MTTKSERLTV LSDAEQEALY GLPDFDDAQR LEYLALTETE LALASSRPGL HAQVYCILQI GYFKAKHAFF RFDWSEVEHD CAFVLSRYFH GESFEHKPIS KHEHYTQREW IADLFGYRPW AAEFLAQLAQ QAAQTVRRDV MPGFIAAELI VWLNEHKIIR PGYTTLQELV SEALSAERRR LAGLLSEVLD ESAKAALGRL LVRDDTLSQL AALKQDAKDF GWRQMARERE KRATLEPLHR IAKALLPKLG VSQQNLLYYA SLANFYTVHD LRNLKADQTY LYLLCYAWVR YRQLSDNLVD AMAYHMKQLE DESSAGAKQS FVAEQVRRQQ DTPQVGRLLS LYIDDSVPDP TPFGDVRQRA YKIMPRDTLQ TTAQRMSVKP VSKLALHWQA VDGLAERIRR HLRPLYVALD LAGTDPGSPW LVALAWAKDV FAKQQRLSQR PLAECPAATL PKRLRPYLLT FDADGKPTGL HADRYEFWLY RQVRKRFQSG ELYLDDSLQH RHFSDELVSL EEKAAVLAQI DIPFLRQPLD AQLDALATEL RSQWLAFNRE LKQGKLTHLE YDKDTQKLTW RKPKGENQKA REKAFYEQLP FCDVADVFRF VNGQCQFLSA LTPLQPRYAK KVADADSLMA VIIAQAMNHG NQVMARTSDI PYHVLESAYQ QYLRHATLHA ANDCISNAIA ALPIFPYYSF DLDALYGAVD GQKFGVERPT VKARHSRKYF GRGKGVVAYT LLCNHVPLNG YLIGAHDYEA HHVFDIWYRN TSDIVPTAIT GDMHSVNKAN FAILHWFGLR FEPRFTDLGD QLKELYSADD PALYDQCLIR PAGRIDRDLI VSEKPNLDQI VATLGLKEMT QGTLIRKLCT YTAPNPTRRA VFEFDKLIRS IYTLRYLRDP QLERNVHRSQ NRIESYHHLR STIAQVGGKK ELTGRTDIEI EISNQCARLI ANAVIFYNSA ILSRLLMKYE VSGNAKAHAL LTQISPAAWR HILLNGHYTF QSDGKMIDLD ELVAGLELG |
||||||||||
1. Oosterkamp MJ, Veuskens T, Plugge CM, Langenhoff AA, Gerritse J, van Berkel WJ, Pieper DH, Junca H, Goodwin LA, Daligault HE, Bruce DC, Detter JC, Tapia R, Han CS, Land ML, Hauser LJ, Smidt H, Stams AJ.
Genome sequences of Alicycliphilus denitrificans strains BC and K601T.. J Bacteriol. 2011 Sep;193(18):5028-9. doi: 10.1128/JB.00365-11. Epub 2011 Jul 8. PubMed ID:
21742888
.
2. Weelink SA, Tan NC, ten Broeke H, van den Kieboom C, van Doesburg W, Langenhoff AA, Gerritse J, Junca H, Stams AJ. Isolation and characterization of Alicycliphilus denitrificans strain BC, which grows on benzene with chlorate as the electron acceptor.. Appl Environ Microbiol. 2008 Nov;74(21):6672-81. doi: 10.1128/AEM.00835-08. Epub 2008 Sep 12. PubMed ID: 18791031 .
2. Weelink SA, Tan NC, ten Broeke H, van den Kieboom C, van Doesburg W, Langenhoff AA, Gerritse J, Junca H, Stams AJ. Isolation and characterization of Alicycliphilus denitrificans strain BC, which grows on benzene with chlorate as the electron acceptor.. Appl Environ Microbiol. 2008 Nov;74(21):6672-81. doi: 10.1128/AEM.00835-08. Epub 2008 Sep 12. PubMed ID: 18791031 .