Mobile element type:
Transposon
Name:
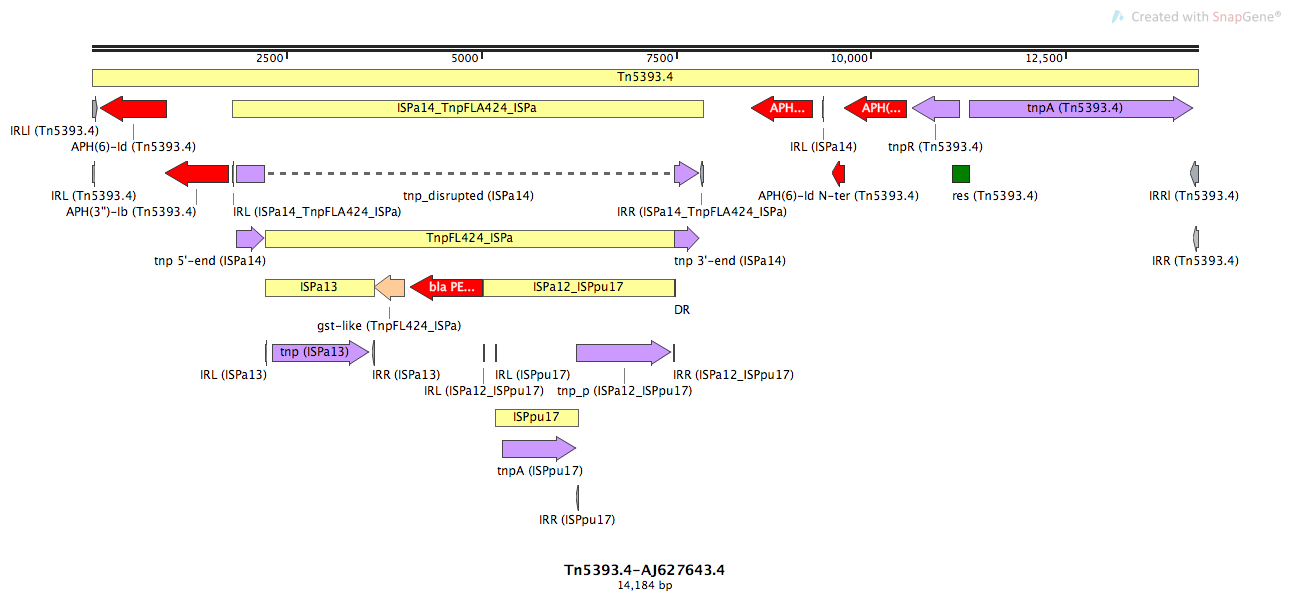
Tn5393.4
Synonyms:
Tn5393d
Accession:
Tn5393.4-AJ627643.4
Family:
Tn3
Group:
Tn163
First isolate:
ND
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Alcaligenes faecalis
Date of Isolation:
2005
Country:
Italy
Molecular Source:
plasmid pFL424
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRLl | 1-81 | + | 81 | GGGGTCGTTT GCGGGAGAGG GCGAAATCCT ACGCTAAGGC TTTGGCCAAC GATATTCTCC GGTAAGATTG ATGTGTTCCC A |
| IRL | 1-40 | + | 40 | GGGGTCGTTT GCGGGAGAGG GCGAAATCCT ACGCTAAGGC |
| IRRl | 14104-14184 | - | 81 | GGGGTCGTTT GCGGGAGGGG GCGGAATCCT ACGCTAAGGC TTTGGCCAGC GATATTCTCC GGTGAGATTG ATGTGTTCCC A |
| IRR | 14145-14184 | - | 40 | GGGGTCGTTT GCGGGAGGGG GCGGAATCCT ACGCTAAGGC |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| res | Tn5393.4 | 11044-11265 | + | 222 | TGTCCCGTTC GACACCTGCG GCGCGCAAGG CGTCGTGCTG CAGGTCGAGA GACTGCGAGC CATCGGCTTT GGAGACGCGG GCATATCCGA TCAGCATGTA TCACAAACGT TGGTTTGAGG CGGCGCTTCG GCCACGATTG CATTGACCTC TGGAAATGTA TCTCAACCAG CTTCATAAAC AAAGCGTCTT GAACGCTATC AGATTTTGAA AAAGGAACAT GT |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| APH(6)-Id (ARO:3002660) | Tn5393.4 | 107-952 | - | Passenger Gene | Antibiotic Resistance |
| APH(3'')-Ib (ARO:3002639) | Tn5393.4 | 943-1746 | - | Passenger Gene | Antibiotic Resistance |
| tnp | ISPa13 | 2321-3580 | + | Transposase | None |
| gst-like | TnpFL424_ISPa | 3621-4009 | - | PassengerGene | Other |
| bla PER-1 (ARO:3002363) | TnpFL424_ISPa | 4083-5009 | - | PassengerGene | Antibiotic Resistance |
| tnpA | ISPpu17 | 5273-6235 | + | Transposase | |
| tnp_p | ISPa12_ISPpu17 | 6219-7451 | + | Transposase | |
| APH(3')-VIa (ARO:3002652) | Tn5393.4 | 8472-9251 | - | Passenger Gene | Antibiotic Resistance |
| APH(3'')-Ib (ARO:3002639) | Tn5393.4 | 9657-10460 | - | Passenger Gene | Antibiotic Resistance |
| tnpR | Tn5393.4 | 10529-11140 | - | Accessory Gene | Resolvase |
| tnpA | Tn5393.4 | 11266-14151 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | APH(6)-Id (ARO:3002660) | APH(6)-Id | Tn5393.4 | 281 | 107-952 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | APH(6) (ARO:3000151) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strB, orfI || strict match to reference sequence for ARO:3002660 (bitscore: 568) | ||||||||||
| Protein Sequence: | MGLMFMPPVF PAHWHVSQPV LIADTFSSLV WKVSLPDGTP AIVKGLKPIE DIADELRGAD YLVWRNGRGA VRLLGRENNL MLLEYAGERM LSHIVAEHGD YQATEIAAEL MAKLYAASEE PLPSALLPIR DRFAALFQRA RDDQNAGCQT DYVHAAIIAD QMMSNASELR GLHGDLHHEN IMFSSRGWLV IDPVGLVGEV GFGAANMFYD PADRDDLCLD PRRIAQMADA FSRALDVDPR RLLDQAYAYG CLSAAWNADG EEEQRDLAIA AAIKQVRQTS Y |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | APH(3'')-Ib (ARO:3002639) | APH(3'')-Ib | Tn5393.4 | 267 | 943-1746 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | APH(3'') (ARO:3000127) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strA, orfH || perfect match to referencesequence for ARO: 3002639 | ||||||||||
| Protein Sequence: | MNRTNIFFGE SHSDWLPVRG GESGDFVFRR GDGHAFAKIA PASRRGELAG ERDRLIWLKG RGVACPEVIN WQEEQEGACL VITAIPGVPA ADLSGADLLK AWPSMGQQLG AVHSLSVDQC PFERRLSRMF GRAVDVVSRN AVNPDFLPDE DKSTPQLDLL ARVERELPVR LDQERTDMVV CHGDPCMPNF MVDPKTLQCT GLIDLGRLGT ADRYADLALM IANAEENWAA PDEAERAFAV LFNVLGIEAP DRERLAFYLR LDPLTWG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnp | Tnp | ISPa13 | 420 | 2321-3580 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | KRMKQQQDTR FSSAFERFFS AAELDALGKQ SGFMKRRRQV TPQKFCMALV SALGAGTTNS IADVHRQFNH MHSTDIKLKP FHKQLVKMAA PEFMREVFEK ALALHLPDMF AIRNKYQDMF KRIILQDGTS FAVHDDLMFY FPGRFNENSP AAVELHVTYD VFKGQPDGVS LTEDTAPERD YLPSPASLSG CLFMADAGYF SKAYIQQLQE ACAYFIMRMC NRVNPMAVCT KTQQMKPLKT WLKELPNAGV LDLDVQWPNG PVYRCVAFAA LDHKKRHVLV CTNLDREQFP PAVVGDFYRV RWQIELLFKE WKSMNNLNKF DTGHATIVET LIWGSLLAAT LKRWLINAAQ KKYNRVLSLF TGAKTAHMWW IPFCFDIAMS RGTDLAKAMN NALAFLAQHC QRQNVKRDQI SGVYKLLNNT |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | gst-like | Gst-like | TnpFL424_ISPa | 129 | 3621-4009 | - |
|---|
| Class: | PassengerGene | ||||||||||
| Subclass: | Other | ||||||||||
| Protein Sequence: | MQLLGSVASP FVRRLRLVLA GQPYQFVALN IFESEGRSVL VQHNPARKVP VLVDGEQVIF DSGVIYRYLA SKLKFKPLSW DQENGLTTIN ACTDSLVELL LCKRSGFDVT EDKLFFNLQH ERIQATLEA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | bla PER-1 (ARO:3002363) | Bla PER-1 | TnpFL424_ISPa | 308 | 4083-5009 | - |
|---|
| Class: | PassengerGene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | PER beta-lactamase (ARO:3000056) | ||||||||||
| Target: | cephalosporin (ARO:0000032)||penem (ARO:3003706)||monobactam (ARO:0000004)||penam (ARO:3000008)||carbapenem (ARO:0000020) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002363 | ||||||||||
| Protein Sequence: | MNVIIKAVVT ASTLLMVSFS SFETSAQSPL LKEQIESIVI GKKATVGVAV WGPDDLEPLL INPFEKFPMQ SVFKLHLAML VLHQVDQGKL DLNQTVIVNR AKVLQNTWAP IMKAYQGDEF SVPVQQLLQY SVSHSDNVAC DLLFELVGGP AALHDYIQSM GIKETAVVAN EAQMHADDQV QYQNWTSMKG AAEILKKFEQ KTQLSETSQA LLWKWMVETT TGPERLKGLL PAGTVVAHKT GTSGIKAGKT AATNDLGIIL LPDGRPLLVA VFVKDSAESS RTNEAIIAQV AQTAYQFELK KLSALSPN |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | ISPpu17 | 320 | 5273-6235 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MTTHYRQLTQ GQRYQIEAGL SAAESQASIA KRVGVHPSTI SREVRRNSTQ NIYKAVSAAH ESDARRAGAR KFCKPATWLS HHLPVWLKHG MSPEQIAQRL KQEQPSRAVS HEWIYRFIAA EQRAGGELYT YLRHRRKRYR KRYGSHDRRG QLRNRVSISE RPAEVESRER LGDWEGDTVH GLGGNLVTLV DRKSGYLSAY PVKRRTRRQV TRAINLMLQG HAAHTLTLDN GREFAGHERI ALRSQCQVFF ADPYSSWQRG TNENTNGLLR QYFPKGSDFS KLTVEAVNRT VARINLRPRK RLGWKTPYEV HTGVSVALMC |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnp_p | Tnp_p | ISPa12_ISPpu17 | 410 | 6219-7451 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | LHLCVEFRQQ LNSLGVQTHM IERFRLITPA KLCLAFVCAL GSGNARTIAD IHRYFNHLHS MSVRLKPFHN QLVKLGTPEF MRQVFEQALA LHLPAMHTFS DAYRGHFKQV LLQDGTSFAV HDGLSLHFPG RFSTHSPAAV ELHVTYDLEK AQPVRVSLSE DTASERDYLP VAQSLRGCLL MADAGYFSKA YIESLQNEAA SFVLRMPASV NPMATCNQTG LCQPLRSWLA VLPKHGELDL DVQWPDGPVY RCVLFASTDH KDKPVCLCTN LDRHTFPAAT VGEWYRLRWQ IELLFKEWKS LNSLNKFNTE YSTIAETLIW GSLLAATLKR WLINGAQQKY RRVLSMFSGA KTANQWWLPF CIKWACSGIT EIAAELGKVF AFLAEHCQRV NVKRDQKSGA FKVLEITYES |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | APH(3')-VIa (ARO:3002652) | APH(3')-VIa | Tn5393.4 | 259 | 8472-9251 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | APH(3') (ARO:3000126) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | loose match to reference sequence for ARO:3002652 (93% identical, bitscore:470)||Synonyms: aphA-6 | ||||||||||
| Protein Sequence: | MELPNIIQQF IGNSVLEPNK IGQSPSDVYS FNRNNETFFL KRSSTLYTET TYSVSREAKM LSWLSDKLKV PELIMTFQDE QFEFMITKAI NAKSISALFL TEQELLAIYK ETLNQLNAVA IIDCPFISSI DHRLKESKFF IDNQLLDEID QDDFEAELWG DHKTYISLWN ELNETRVEER LVFSHGDITD SNIFIDKSGE IYFLDLGRAG LADEFVDISF VERCLREDVS EETAKIFLKH LKNDMPDKRN YFLKLDELN |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | APH(3'')-Ib (ARO:3002639) | APH(3'')-Ib | Tn5393.4 | 267 | 9657-10460 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | APH(3'') (ARO:3000127) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strA, orfH || perfect match to referencesequence for ARO: 3002639 | ||||||||||
| Protein Sequence: | MNRTNIFFGE SHSDWLPVRG GESGDFVFRR GDGHAFAKIA PASRRGELAG ERDRLIWLKG RGVACPEVIN WQEEQEGACL VITAIPGVPA ADLSGADLLK AWPSMGQQLG AVHSLSVDQC PFERRLSRMF GRAVDVVSRN AVNPDFLPDE DKSTPQLDLL ARVERELPVR LDQERTDMVV CHGDPCMPNF MVDPKTLQCT GLIDLGRLGT ADRYADLALM IANAEENWAA PDEAERAFAV LFNVLGIEAP DRERLAFYLR LDPLTWG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | Tn5393.4 | 204 | 10529-11140 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Function: | recombinaseactivity (GO:0000150) | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MLIGYARVSK ADGSQSLDLQ HDALRAAGVE RDNIYDDLAS GGRDDRPGLT ACLKSLRDGD VLVVWKLDRL GRSLAHLVNT VKELSDRKIG LRVLTGKGAQ IDTTTASGRM VFGIFATLAE FERDLIRERT MAGLASARAR GRKGGRKFAL TKAQVRLAQA AMAQRDTSVS DLCKELGIER VTLYRYVGPK GELRDHGKHV LGLT |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn5393.4 | 961 | 11266-14151 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase activity (GO:0004803) | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MPRRVTLTDR QKDALLRLPT SQTDLLKHYT LSDEDLGHIR LRRRAHNRFG FALQLCVLRY PGRVLAPGEL IPAEVIEFIG AQLGLGADDL VDYAAREETR HEHLAELRGL YGFRTFSGRG ASELKEWLFR EAEMAVSNED IARRFVAECR RTRTVLPATS TIERLCAAAL VDAERRIETR IASRLPMSIR EQLLALLEET ADDRVTRFVW LRQFEPGSNS SSANRLLDRL EYLQRIDLPE DLLAGVPAHR VTRLRRQGER YYADGMRDLP EDRRLAILAV CVSEWQAMLA DAVVETHDRI VGRLYRASER ICHAKVADEA GVVRDTLKSF AEIGGALVDA QDDGQPLGDV IASGSGWDGL KTLVAMATRL TATMADDPLN HVLDGYHRFR RYAPRMLRLL DLRAAPVALP LLEAVTALRT GLNDAAMTSF LRPSSKWHRH LRAQRAGDAR LWEIAVLFHL RDAFRSGDVW LTRSRRYGDL KHALVPAQSI AEGGRLAVPL RPEEWLADRQ ARLDMRLREL GRAARAGTIP GGSIENGVLH IEKLEAAAPT GAEDLVLDLY KQIPPTRITD LLLEVDAATG FTEAFTHLRT GAPCADRIGL MNVILAEGIN LGLRKMADAT NTHTFWELIR IGRWHVEGEA YDRALAMVVE AQAALPMARF WGMGTSASSD GQFFVATEQG EAMNLVNAKY GNTPGLKAYS HVSDQYAPFA TQVIPATASE APYILDGLLM NDAGRHIREQ FTDTGGFTDH VFAACAILGY RFAPRIRDLP SKRLYAFNPS AAPAHLRALI GGKVNQAMIE RNWPDILRIA ATIAAGTVAP SQILRKLASY PRQNELATAL REVGRVERTL FMIDWILDAE LQRRAQIGLN KGEAHHALKR AISFHRRGEI RDRSAEGQHY RIAGMNLLAA IIIFWNTMKL GEVVANQKRD GKLLSPDLLA HVSPLGWEHI NLTGEYRWPK P |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| ISPa14_TnpFLA424_ISPa-AJ6276431 | ISPa14_TnpFLA424_ISPa | insertion sequence | 1803-7844 | + | 6042 |
| TnpFL424_ISPa-AJ627643 | TnpFL424_ISPa | transposon | 2234-7479 | + | 5246 |
| ISPa13-AJ627643 | ISPa13 | insertion sequence | 2234-3620 | + | 1387 |
| ISPa12_ISPpu17-AJ627643.4 | ISPa12_ISPpu17 | insertion sequence | 5023-7479 | + | 2457 |
| ISPpu17-DQ174113.1 | ISPpu17 | insertion sequence | 5177-6242 | + | 1066 |
| Name | Associated TE | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|---|
| IRL | ISPa14_TnpFLA424_ISPa | 1803-1829 | + | 27 | GGTGGTGTTT CAAAAAGTAT GCTGAAA |
| IRL | ISPa13 | 2234-2254 | + | 21 | TCATAGGTAT GATCTTTAGG A |
| IRR | ISPa13 | 3600-3620 | - | 21 | TCATACGTAT GACCTTAACG A |
| IRL | ISPa12_ISPpu17 | 5023-5043 | + | 21 | TCATACGTAT GATCTTTAGG A |
| IRL | ISPpu17 | 5177-5198 | + | 22 | CCTGAATTCA TAGAATAAGC GC |
| IRR | ISPpu17 | 6221-6242 | - | 22 | CCTGAATTCA ACACATAAGT GC |
| IRR | ISPa12_ISPpu17 | 7459-7479 | - | 21 | TCATACGTAT GTCTTTAGCG A |
| IRR | ISPa14_TnpFLA424_ISPa | 7818-7844 | - | 27 | GGTGGTGTTT CAAAAAGTAT GCTGACA |
1. Mantengoli E, Rossolini GM.
Tn5393d, a complex Tn5393 derivative carrying the PER-1 extended-spectrum beta-lactamase gene and other resistance determinants.. Antimicrob Agents Chemother. 2005 Aug;49(8):3289-96. doi: 10.1128/AAC.49.8.3289-3296.2005. PubMed ID:
16048938
.