Mobile element type:
Transposon
Name:
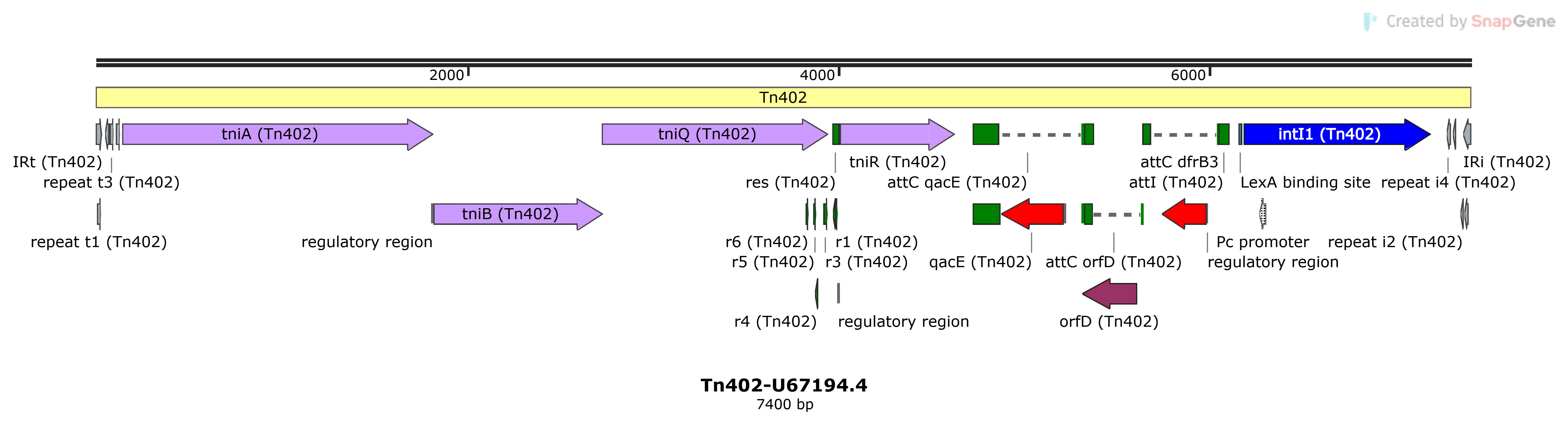
Tn402
Synonyms:
Tn5090
Accession:
Tn402-U67194.4
Family:
Tn402
Group:
First isolate:
Yes
Partial:
ND
Evidence of transposition:
Yes
Host Organism:
Klebsiella aerogenes(capsular type 37)
Date of Isolation:
1977
Country:
United kingdom
Molecular Source:
plasmid R375
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| None | 1-33 | + | 33 | TGTCATTTTC AGAAGACGAC TGCACCAGTT GAT |
| repeat t1 | 9-27 | + | 19 | TCAGAAGACG ACTGCACCA |
| repeat t2 | 49-67 | - | 19 | TCAGGAGCTG GCTGCACAA |
| repeat t3 | 78-97 | + | 20 | TCAGAAGTGA TCTGCACCAA |
| repeat t4 | 110-128 | + | 19 | TCAATACTCG TGTGCACCA |
| repeat i4 | 7281-7299 | - | 19 | TCAGCGGACG CAGGGAGGA |
| repeat i3 | 7309-7327 | - | 19 | TCAGGCAACG ACGGGCTGC |
| repeat i2 | 7351-7369 | - | 19 | TCAGAAGCCG ACTGCACTA |
| None | 7368-7400 | - | 33 | TGTCGTTTTC AGAAGACGGC TGCACTGAAC GTC |
| repeat i1 | 7374-7392 | - | 19 | TCAGAAGACG GCTGCACTG |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| r6 | Tn402 | 3824-3837 | + | 14 | TTGGAACGGT TGCG |
| r5 | Tn402 | 3870-3883 | + | 14 | AATTTCTGTC ACAT |
| r4 | Tn402 | 3875-3888 | - | 14 | AGGTAATGTG ACAG |
| r3 | Tn402 | 3926-3939 | + | 14 | GGGTTAAGTG ACAA |
| res | Tn402 | 3967-4001 | + | 35 | ACACTGTCAC ATAATCGAAC GTATACGTGA CGGGT |
| r2 | Tn402 | 3970-3983 | - | 14 | CGATTATGTG ACAG |
| r1 | Tn402 | 3986-3999 | + | 14 | CGTATACGTG ACGG |
| attC qacE | Tn402 | 4728-4868 , 5315-5320 | + | 147 | GTTCAACGCC GAGTTCAGCG GCAGTTTTTA AGTTGTGATT TTATGGAATA CTTTTGCGCA GCAAAACCAT AAAGCCGCGA CTTAAAAACT GTCCAGCGCA GGCACGAAGT GCTGGAGCGG TGCTGCAACG ACTTGTTAGA TATCTAA |
| attC qacE core | Tn402 | 4728-4868 | + | 141 | GTTCAACGCC GAGTTCAGCG GCAGTTTTTA AGTTGTGATT TTATGGAATA CTTTTGCGCA GCAAAACCAT AAAGCCGCGA CTTAAAAACT GTCCAGCGCA GGCACGAAGT GCTGGAGCGG TGCTGCAACG ACTTGTTAGA T |
| attC orfD | Tn402 | 5326-5373 , 5635-5641 | + | 55 | GAGGTAAGCC GACCGCAGAA TGCGGGTCGG CTTGACCGAA ATGTTAGATA CTAAC |
| attC orfD core | Tn402 | 5326-5373 | + | 48 | GAGGTAAGCC GACCGCAGAA TGCGGGTCGG CTTGACCGAA ATGTTAGA |
| attI | Tn402 | 6049-6104 | + | 56 | CTTTGTTTTA GGGCGACTGC CCTGCTGCGT AACATCGTTG CTGCTCCATA ACATCA |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tniA | Tn402 | 142-1821 | + | Transposase | |
| tniB | Tn402 | 1824-2732 | + | Accessory Gene | |
| tniQ | Tn402 | 2729-3946 | + | Accessory Gene | Target Site Selection |
| tniR | Tn402 | 4008-4631 | + | Accessory Gene | Resolvase |
| qacE (ARO:3005009) | Tn402 | 4880-5212 | - | Passenger Gene | Antibiotic Resistance |
| dfrB3 (ARO:3003022) | Tn402 | 5744-5980 | - | Passenger Gene | Antibiotic Resistance |
| intI1 | Tn402 | 6185-7198 | + | Integron Integrase | Class 1 |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA | TniA | Tn402 | 559 | 142-1821 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | can be extended upstream by 12 amino acids| identical to tniA (Tn1721 and In2)| 25% amino acid sequence identity to TnsB from Tn7 | ||||||||||
| Protein Sequence: | MATDTPRIPE QGVATLPDEA WERARRRAEI ISPLAQSETV GHEAADMAAQ ALGLSRRQVY VLIRRARQGS GLVTDLVPGQ SGGGKGKGRL PEPVERVIHE LLQKRFLTKQ KRSLAAFHRE VTQVCKAQKL RVPARNTVAL RIASLDPRKV IRRREGQDAA RDLQGVGGEP PAVTAPLEQV QIDHTVIDLI VVDDRDRQPI GRPYLTLAID VFTRCVLGMV VTLEAPSAVS VGLCLVHVAC DKRPWLEGLN VEMDWQMSGK PLLLYLDNAA EFKSEALRRG CEQHGIRLDY RPLGQPHYGG IVERIIGTAM QMIHDELPGT TFSNPDQRGD YDSENKAALT LRELERWLTL AVGTYHGSVH NGLLQPPAAR WAEAVARVGV PAVVTRATSF LVDFLPILRR TLTRTGFVID HIHYYADALK PWIARRERWP SFLIRRDPRD ISRIWVLEPE GQHYLEIPYR TLSHPAVTLW EQRQALAKLR QQGREQVDES ALFRMIGQMR EIVTSAQKAT RKARRDADRR QHLKTSARPD KPVPPDTDIA DPQADNLPPA KPFDQIEEW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB | TniB | Tn402 | 302 | 1824-2732 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Sequence Family: | ATP binding protein? | ||||||||||
| Comment: | identical to tniB (Tn1721)| similar function to Tn7 tnsC and MuB | ||||||||||
| Protein Sequence: | MDEYPIIDLS HLLPAAQGLA RLPADERIQR LRADRWIGYP RAVEALNRLE ALYAWPNKQR MPNLLLVGPT NNGKSMIVEK FRRTHPASSD ADQEHIPVLV VQMPSEPSVI RFYVALLAAM GAPLRPRPRL PEMEQLALAL LRKVGVRMLV IDELHNVLAG NSVNRREFLN LLRFLGNELR IPLVGVGTRD AYLAIRSDDQ LENRFEPMML PVWEANDDCC SLLASFAASL PLRRPSPIAT LDMARYLLTR SEGTIGELAH LLMAAAIVAV ESGEEAINHR TLSMAVYTGP SERRRQFERE LM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniQ | TniQ | Tn402 | 405 | 2729-3946 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Target Site Selection | ||||||||||
| Comment: | identical to tniQ (Tn1721)|similar function to Tn7 tnsD? | ||||||||||
| Protein Sequence: | MKPAPRWPLH PAPKEGEALS SWLNRVALCY HMEEPDLLEH DLGHGQVDDL DTAPPLSLLA LLSQRSGIEL DRLRCMSFAG WVPWLLDSLD DQIPDALETY AFQLSVLLPR LRRKTRSITS WRAWLPSQPI NRACPLCLSD PENQAVLLAW KLPLMLSCPL HGCWLESYWG VPGRFLGWEN ADAEPRTASD AIAAMDQRTW QALTTGHVEL PRRRIHAGLW FRLLRTLLDE LNTPLSACGT CAGYPRQVWE GCGHPLRAGQ SLWRPYETLN PIVRLQMLEA AATAISLIEV RDISPPGEQA KLFWSEPQTG FTSGLPTKAP KPEPINHWQR AVQAIDEAII EARHNPETAR SLFALASYGR RDPASLERLR ATFVKEGIPP EFLSHYLPDA PFACLKQNDG LSDKF |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniR | TniR | Tn402 | 207 | 4008-4631 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Comment: | resolution of cointegrates || Protein: ACE81792.1 || identical to tniR (Tn1721) | ||||||||||
| Protein Sequence: | MLIGYMRVSK ADGSQSTNLQ RDALIAAGVS LAHLYEDLAS GRRDDRPGLA ACLKALREGD TLIVWKLDRL GRDLRHLINT VHDLTARSVG LKVLTGHGAA VDTTTAAGKL VFGIFAALAE FERELISERT VAGLISARAR GRKGGRPFKM TAAKLRLAMA SMGQPETKVG DLCEELGITR QTLYRHVSPK GELRPDGVKL LSLGSAA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacE (ARO:3005009) | QacE | Tn402 | 110 | 4880-5212 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | SMR export pump||strict match to reference sequence for ARO:3005009 (bitscore:204) | ||||||||||
| Protein Sequence: | MKGWLFLVIA IVGEVIATSA LKSSEGFTKL APSAVVIIGY GIAFYFLSLV LKSIPVGVAY AVWSGLGVVI ITAIAWLLHG QKLDAWGFVG MGLIVSGVVV LNLLSKASAH |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | dfrB3 (ARO:3003022) | DfrB3 | Tn402 | 78 | 5744-5980 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | trimethoprim resistant dihydrofolate reductase dfr (ARO:3001218) | ||||||||||
| Target: | diaminopyrimidine antibiotic (ARO:3000171) | ||||||||||
| Comment: | 100% identity to reference sequence ARO:3003022 in Klebsiella oxytoca (bitscore: 163) | ||||||||||
| Protein Sequence: | MDQHNNGVST LVAGQFALPS HATFGLGDRV RKKSGAAWQG QVVGWYCTKL TPEGYAVESE SHPGSVQIYP VAALERVA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI1 | IntI1 | Tn402 | 337 | 6185-7198 | + |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 1 | ||||||||||
| Sequence Family: | Class 1 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MKTATAPLPP LRSVKVLDQL RERIRYLHYS LRTEQAYVNW VRAFIRFHGV RHPATLGSSE VEAFLSWLAN ERKVSVSTHR QALAALLFFY GKVLCTDLPW LQEIGRPRPS RRLPVVLTPD EVVRILGFLE GEHRLFAQLL YGTGMRISEG LQLRVKDLDF DHGTIIVREG KGSKDRALML PESLAPSLRE QLSRARAWWL KDQAEGRSGV ALPDALERKY PRAGHSWPWF WVFAQHTHST DPRSGVVRRH HMYDQTFQRA FKRAVEQAGI TKPATPHTLR HSFATALLRS GYDIRTVQDL LGHSDVSTTM IYTHVLKVGG AGVRSPLDAL PPLTSER |
||||||||||
1. Shapiro JA, Sporn P.
Tn402: a new transposable element determining trimethoprim resistance that inserts in bacteriophage lambda.. J Bacteriol. 1977 Mar;129(3):1632-5. doi: 10.1128/jb.129.3.1632-1635.1977. PubMed ID:
321437
.
2. Ferrante AA, Lessie TG. Nucleotide sequence of IS402 from Pseudomonas cepacia.. Gene. 1991 Jun 15;102(1):143-4. doi: 10.1016/0378-1119(91)90555-p. PubMed ID: 1650732 .
3. Marchiaro P, Viale AM, Ballerini V, Rossignol G, Vila AJ, Limansky A. First report of a Tn402-like class 1 integron carrying blaVIM-2 in Pseudomonas putida from Argentina.. J Infect Dev Ctries. 2010 Jun 30;4(6):412-6. PubMed ID: 20601796 .
4. Sajjad A, Holley MP, Labbate M, Stokes HW, Gillings MR. Preclinical class 1 integron with a complete Tn402-like transposition module.. Appl Environ Microbiol. 2011 Jan;77(1):335-7. doi: 10.1128/AEM.02142-10. Epub 2010 Oct 29. PubMed ID: 21037292 .
5. Smith CA, Thomas CM. Comparison of the nucleotide sequences of the vegetative replication origins of broad host range IncP plasmids R751 and RK2 reveals conserved features of probable functional importance.. Nucleic Acids Res. 1985 Jan 25;13(2):557-72. doi: 10.1093/nar/13.2.557. PubMed ID: 4000925 .
2. Ferrante AA, Lessie TG. Nucleotide sequence of IS402 from Pseudomonas cepacia.. Gene. 1991 Jun 15;102(1):143-4. doi: 10.1016/0378-1119(91)90555-p. PubMed ID: 1650732 .
3. Marchiaro P, Viale AM, Ballerini V, Rossignol G, Vila AJ, Limansky A. First report of a Tn402-like class 1 integron carrying blaVIM-2 in Pseudomonas putida from Argentina.. J Infect Dev Ctries. 2010 Jun 30;4(6):412-6. PubMed ID: 20601796 .
4. Sajjad A, Holley MP, Labbate M, Stokes HW, Gillings MR. Preclinical class 1 integron with a complete Tn402-like transposition module.. Appl Environ Microbiol. 2011 Jan;77(1):335-7. doi: 10.1128/AEM.02142-10. Epub 2010 Oct 29. PubMed ID: 21037292 .
5. Smith CA, Thomas CM. Comparison of the nucleotide sequences of the vegetative replication origins of broad host range IncP plasmids R751 and RK2 reveals conserved features of probable functional importance.. Nucleic Acids Res. 1985 Jan 25;13(2):557-72. doi: 10.1093/nar/13.2.557. PubMed ID: 4000925 .