Mobile element type:
Transposon
Name:
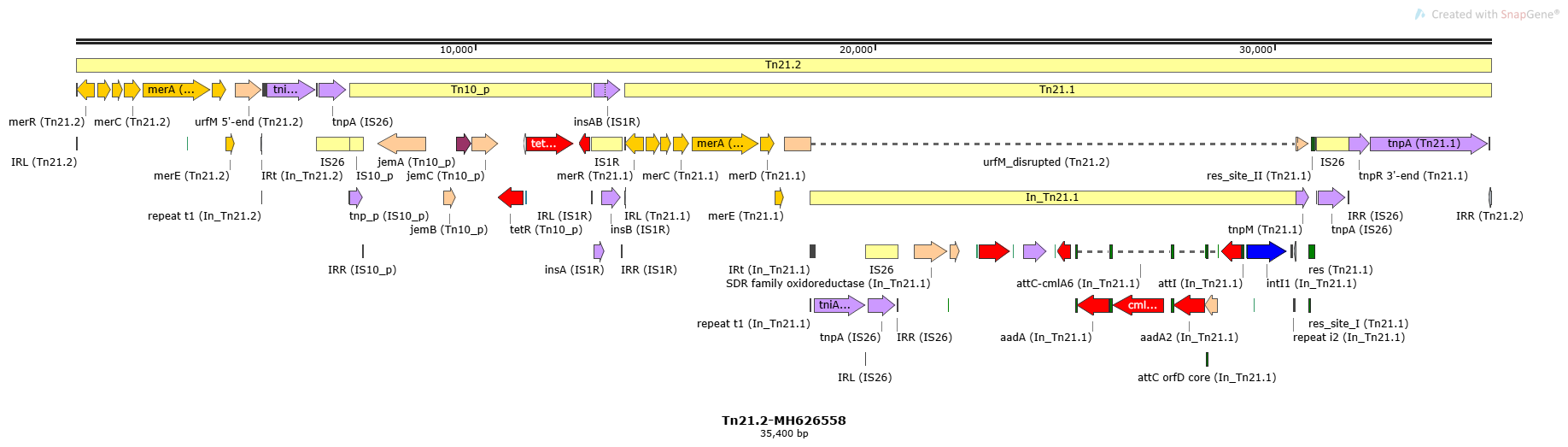
Tn21.2
Synonyms:
Accession:
Tn21.2-MH626558
Family:
Tn3
Group:
Tn21
First isolate:
ND
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Salmonella enterica subsp. enterica serovar Typhimurium
Date of Isolation:
Country:
Molecular Source:
plasmid pST1007-1B
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGGCACCT CAGAAAACGG AAAATAAAGC ACGCTAAG |
| repeat i4 | 13742-13760 | + | 19 | TCAGAAAACG GAAAATAAA |
| IRR | 35360-35400 | - | 41 | GGGGTCGTCT CAGAAAACGG AAAATAAAGC ACGCTAAGCC G |
| repeat i4 | 35373-35391 | - | 19 | TCAGAAAACG GAAAATAAA |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| attC-cmlA6 | In_Tn21.1 | 25873-25936 , 27416-27421 | + | 70 | CGCCTGAGCT CAGCCGACCG AAACCGCGTA GCGGTTTTGG GTCGGCTGCA GCGATTTGTT GGGCGCCCAA |
| attC-cmlA6 5'-end | In_Tn21.1 | 25873-25936 | + | 64 | CGCCTGAGCT CAGCCGACCG AAACCGCGTA GCGGTTTTGG GTCGGCTGCA GCGATTTGTT GGGC |
| attC cmlA6 core | In_Tn21.1 | 25873-25936 | + | 64 | CGCCTGAGCT CAGCCGACCG AAACCGCGTA GCGGTTTTGG GTCGGCTGCA GCGATTTGTT GGGC |
| attC-cmlA6 3'-end | In_Tn21.1 | 27416-27421 | + | 6 | GCCCAA |
| attC-aadA2 | In_Tn21.1 | 27422-27475 , 28272-28277 | + | 60 | CGCCGGAGTT AAGCCGCCGC GCGTAGCGCG GTCGGCTTGA ACGAATTGTT AGACGTCTAA |
| attC-aadA2 5'-end | In_Tn21.1 | 27422-27475 | + | 54 | CGCCGGAGTT AAGCCGCCGC GCGTAGCGCG GTCGGCTTGA ACGAATTGTT AGAC |
| attC aadA3 core | In_Tn21.1 | 27422-27475 | + | 54 | CGCCGGAGTT AAGCCGCCGC GCGTAGCGCG GTCGGCTTGA ACGAATTGTT AGAC |
| attC-aadA2 3'-end | In_Tn21.1 | 28272-28277 | + | 6 | GTCTAA |
| attC orfD core | In_Tn21.1 | 28283-28330 | + | 48 | GAGGTAAGCC GACCGCAGAA TGCGGGTCGG CTTGACCGAA ATGTTAGA |
| attI | In_Tn21.1 | 29182-29237 | + | 56 | CTTTGTTTTA GGGCGACTGC CCTGCTGCGT AACATCGTTG CTGCTCCATA ACATCA |
| res | Tn21.1 | 30869-30999 | + | 131 | GCCGCCGTCA GGTTGAGGCA TACCCTAACC TGATGTCAGA TGCCATGTGT AAATTGCGTC AGGATAGGAT TGAATTTTGA ATTTATTGAC ATATCTCGTT GAAGGTCATA GAGTCTTCCC TGACATTTTG C |
| res_site_I | Tn21.1 | 30869-30907 | + | 39 | GCCGCCGTCA GGTTGAGGCA TACCCTAACC TGATGTCAG |
| res_site_II | Tn21.1 | 30921-30964 | + | 44 | ATTGCGTCAG GATAGGATTG AATTTTGAAT TTATTGACAT ATCT |
| res_site_III | Tn21.1 | 30968-30999 | + | 32 | TGAAGGTCAT AGAGTCTTCC CTGACATTTT GC |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| merR | Tn21.2 | 34-468 | - | Passenger Gene | Heavy Metal Resistance |
| merT | Tn21.2 | 540-890 | + | Passenger Gene | Heavy Metal Resistance |
| merP | Tn21.2 | 904-1179 | + | Passenger Gene | Heavy Metal Resistance |
| merC | Tn21.2 | 1215-1637 | + | Passenger Gene | Heavy Metal Resistance |
| merA | Tn21.2 | 1689-3383 | + | Passenger Gene | Heavy Metal Resistance |
| merD | Tn21.2 | 3401-3763 | + | Passenger Gene | Heavy Metal Resistance |
| merE | Tn21.2 | 3760-3996 | + | Passenger Gene | Heavy Metal Resistance |
| urfM 5'-end | Tn21.2 | 3993-4663 | + | Passenger Gene | Other |
| tniA_p | Tn21.2 | 4775-6026 | + | Transposase | |
| tnpA | IS26 | 6090-6794 | + | Transposase | |
| tnp_p | IS10_p | 6848-7186 | + | Transposase | |
| jemA | Tn10_p | 7552-8757 | - | Passenger Gene | Other |
| jemB | Tn10_p | 9201-9521 | + | Passenger Gene | Other |
| CPT07_26605 | Tn10_p | 9514-9900 | + | Passenger Gene | Hypothetical |
| jemC | Tn10_p | 9908-10594 | + | Passenger Gene | Other |
| tetR (ARO:3003479) | Tn10_p | 10572-11198 | - | Passenger Gene | Antibiotic Resistance |
| tet(B) (ARO:3000166) | Tn10_p | 11277-12482 | + | Passenger Gene | Antibiotic Resistance |
| tetC_p | Tn10_p | 12595-12849 | - | Passenger Gene | Antibiotic Resistance |
| insAB | IS1R | 12962-13659 | + | Transposase | |
| insA | IS1R | 12962-13237 | + | Accessory Gene | |
| insB | IS1R | 13156-13659 | + | Transposase | |
| merR | Tn21.1 | 13766-14200 | - | Passenger Gene | Heavy Metal Resistance |
| merT | Tn21.1 | 14272-14622 | + | Passenger Gene | Heavy Metal Resistance |
| merP | Tn21.1 | 14636-14911 | + | Passenger Gene | Heavy Metal Resistance |
| merC | Tn21.1 | 14947-15369 | + | Passenger Gene | Heavy Metal Resistance |
| merA | Tn21.1 | 15421-17115 | + | Passenger Gene | Heavy Metal Resistance |
| merD | Tn21.1 | 17133-17495 | + | Passenger Gene | Heavy Metal Resistance |
| merE | Tn21.1 | 17492-17728 | + | Passenger Gene | Heavy Metal Resistance |
| urfM 5'-end | Tn21.2 | 17725-18395 | + | Passenger Gene | Other |
| tniA_p | In_Tn21.1 | 18471-19775 | + | Transposase | |
| tnpA | IS26 | 19822-20526 | + | Transposase | |
| SDR family oxidoreductase | In_Tn21.1 | 20984-21847 | + | Passenger Gene | Other |
| GrpB domain protein | In_Tn21.1 | 21885-22130 | + | Passenger Gene | Other |
| sul3 (ARO:3000413) | In_Tn21.1 | 22599-23390 | + | Passenger Gene | Antibiotic Resistance |
| tnp IS256 family | In_Tn21.1 | 23715-24323 | + | Transposase | |
| qacL (ARO:3005098) | In_Tn21.1 | 24570-24902 | - | Passenger Gene | Antibiotic Resistance |
| aadA (ARO:3002601) | In_Tn21.1 | 25072-25863 | - | Passenger Gene | Antibiotic Resistance |
| cmlA6 (ARO:3002696) | In_Tn21.1 | 25956-27215 | - | Passenger Gene | Antibiotic Resistance |
| aadA2 (ARO:3002602) | In_Tn21.1 | 27477-28256 | - | Passenger Gene | Antibiotic Resistance |
| DUF1010 family protein | In_Tn21.1 | 28274-28564 | - | Passenger Gene | Other |
| dfrA12 (ARO:3002858) | In_Tn21.1 | 28676-29173 | - | Passenger Gene | Antibiotic Resistance |
| intI1 | In_Tn21.1 | 29318-30331 | + | Integron Integrase | Class 1 |
| tnpM | Tn21.1 | 30534-30884 | + | Accessory Gene | Inhibitor |
| tnpR 5'-end | Tn21.1 | 31010-31032 | + | Accessory Gene | Resolvase |
| tnpA | IS26 | 31104-31808 | + | Transposase | |
| tnpR 3'-end | Tn21.1 | 31861-32398 | + | Accessory Gene | Resolvase |
| tnpA | Tn21.1 | 32401-35367 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merR | MerR | Tn21.2 | 144 | 34-468 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | activator-repressor of mer operon | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MENNLENLTI GVFAKAAGVN VETIRFYQRK GLLREPDKPY GSIRRYGEAD VVRVKFVKSA QRLGFSLDEI AELLRLDDGT HCEEASSLAE HKLKDVREKM ADLARMETVL SELVCACHAR KGNVSCPLIA SLQGEAGLAR SAMP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merT | MerT | Tn21.2 | 116 | 540-890 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | cytosolic mercuric ion transport protein | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSEPQNGRGA LFAGGLAAIL ASTCCLGPLV LVALGFSGAW IGNLTVLEPY RPLFIGAALV ALFFAWKRIY RPVQACKPGE VCAIPQVRAT YKLIFWIVAV LVLVALGFPY VVPFFY |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merP | MerP | Tn21.2 | 91 | 904-1179 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MKKLFASLAL AAAVAPVWAA TQTVTLAVPG MTCAACPITV KKALSKVEGV SKVDVGFEKR EAVVTFDDTK ASVQKLTKAT ADAGYPSSVK Q |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merC | MerC | Tn21.2 | 140 | 1215-1637 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | transmembrane protein mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MGLMTRIADK TGALGSVVSA MGCAACFPAL ASFGAAIGLG FLSQYEGLFI SRLLPLFAAL AFLANALGWF SHRQWLRSLL GMIGPAIVFA ATVWLLGNWW TANLMYVGLA LMIGVSIWDF VSPAHRRCGP DGCELPAKRL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merA | MerA | Tn21.2 | 564 | 1689-3383 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercuric ion reductase | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSTLKITGMT CDSCAVHVKD ALEKVPGVQS ADVSYAKGSA KLAIEVGTSP DALTAAVAGL GYRATLADAP SVSTPGGLLD KMRDLLGRND KTGSSGALHI AVIGSGGAAM AAALKAVEQG ARVTLIERGT IGGTCVNVGC VPSKIMIRAA HIAHLRRESP FDGGIAATTP TIQRTALLAQ QQARVDELRH AKYEGILEGN PAITVLHGSA RFKDNRNLIV QLNDGGERVV AFDRCLIATG ASPAVPPIPG LKDTPYWTST EALVSETIPK RLAVIGSSVV ALELAQAFAR LGAKVTILAR STLFFREDPA IGEAVTAAFR MEGIEVREHT QASQVAYING EGDGEFVLTT AHGELRADKL LVATGRAPNT RKLALDATGV TLTPQGAIVI DPGMRTSVEH IYAAGDCTDQ PQFVYVAAAA GTRAAINMTG GDAALNLTAM PAVVFTDPQV ATVGYSEAEA HHDGIKTDSR TLTLDNVPRA LANFDTRGFI KLVVEEGSGR LIGVQAVAPE AGELIQTAAL AIRNRMTVQE LADQLFPYLT MVEGLKLAAQ TFNKDVKQLS CCAG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merD | MerD | Tn21.2 | 120 | 3401-3763 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | secondary regulatory protein | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSAYTVSQLA HNAGVSVHIV RDYLVRGLLR PVACTTGGYG VFDDAALQRL CFVRAAFEAG IGLDALARLC RALDAADGAQ AAAQLAVLRQ LVERRRAALA HLDAQLASMP AERAHEEALP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merE | MerE | Tn21.2 | 78 | 3760-3996 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Comment: | similar to urf-1 in pKLH2 (GenBank AF213017), pKLH272 (Genbank Y08992), pMER610 (GenBank Y08993), pKLH210 (GenBank Y10102), Tn5036 (Genbank Y09025), orf1 in Tn501 (GenBank Z00027), and urf-1 in Tn5041 (GenBank X98999) | ||||||||||
| Protein Sequence: | MNAPDKLPPE TRQPVSGYLW GALAVLTCPC HLPILAAVLA GTTAGAFLGE HWGVAALALT GLFVLAVTRL LRAFRGGS |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | urfM 5'-end | N/A | Tn21.2 | 223 | 3993-4663 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Comment: | urfMORF interrupted by insertion of In2 | ||||||||||
| Protein Sequence: | MTSSQPAGWT AAELAQAAAR GQLDLHYQPL VDLRDHRIAG AEALMRWRHP RLGLLPPGQF LPLAESFGLM PEIGAWVLGE ACRQMHKWQG PAWQPFRLAI NVSASQVGPT FDDEVKRVLA DMALPAELLE IELTESVAFG NPALFASFDA LRAIGVRFAA DDFGTGYSCL QHLKCCPITT LKIDQSFVAR LPDDARDQTI VRAVIQLAHG LGMDVIFRRR LHQ |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA_p | TniA_p | Tn21.2 | 417 | 4775-6026 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | Contains the first 417 amino acids of tniA (In2)|| can be extended upstream by 12 amino acids|| truncated by insertion of IS26 | ||||||||||
| Protein Sequence: | MATDTPRIPE QGVATLPDEA WERARRRAEI ISPLAQSETV GHEAADMAAQ ALGLSRRQVY VLIRRARQGS GLVTDLVPGQ SGGGKGKGRL PEPVERVIHE LLQKRFLTKQ KRSLAAFHRE VTQVCKAQKL RVPARNTVAL RIASLDPRKV IRRREGQDAA RDLQGVGGEP PAVTAPLEQV QIDHTVIDLI VVDDRDRQPI GRPYLTLAID VFTRCVLGMV VTLEAPSAVS VGLCLVHVAC DKRPWLEGLN VEMDWQMSGK PLLLYLDNAA EFKSEALRRG CEQHGIRLDY RPLGQPHYGG IVERIIGTAM QMIHDELPGT TFSNPDQRGD YDSENKAALT LRELERWLTL AVGTYHGSVH NGLLQPPAAR WAEAVARVGV PAVVTRATSF LVDFLPILRR TLTRTGFVID HIHYYAD |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 6090-6794 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnp_p | N/A | IS10_p | 112 | 6848-7186 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | highly similar to C terminal 112 amino acids of tnp (IS10R) | ||||||||||
| Protein Sequence: | IEETFRDLKS PAYGLGLRHS RTSSSERFDI MLLIALMLQL TCWLAGVHAQ KQGWDKHFQA NTVRNRNVLS TVRLGMEVLR HSGYTITRED SLVAATLLTQ NLFTHGYVLG KL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | jemA | JemA | Tn10_p | 401 | 7552-8757 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Function: | glutamate:sodium symporter activity (GO:0015501) | ||||||||||
| Comment: | sodium-dependent glutamate permease | ||||||||||
| Protein Sequence: | MILDASYTLL VACIALLIGM FVVKFTPFLQ KNHIPEAVVG GFIVAIVLLI IDKTSGYSFT FDASLQSLLM LTFFSSIGLS SDFSRLIKGG KPLVLLTIAV TILIAIQNTV GMSMAVMMNE SPFIGLIAGS ITLTGGHGNA GAWGPILADK YGVTGAVELA MACATLGLVL GGLVGGPVAR HLLKKVSIPK TTEQERDTIV EAFEQPSVKR KINANNVIET ISMLIICIVV GGYISALFKD TFLQLPTFVW CLFVGIIIRN TLTHVFKHEV FEPTVDVLGS VALSLFLAMA LMSLKFGQLA SMAGPVLIII AVQTVVMVLF ACFVTFKMMG KDYDAVVISA GHCGFGMGAT PTAIANMQTV TKAFGPSHKA FLVVPMVGAF IVDISNSILI KIFIEIGTYF T |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | jemB | JemB | Tn10_p | 106 | 9201-9521 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | Antibiotic biosynthesis monooxygenase (Pfam:PF03992) | ||||||||||
| Protein Sequence: | MIAVIFEVQI QPDQQTRYLT LAEELRPLLS HVAGFISIER FQSLATEGKM LSLSWWENEY AVLQWKNHVL HAKAQQEGRE SIFDFYKISI AHITREYSFK KDKDNV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | CPT07_26605 | CPT07_26605 | Tn10_p | 128 | 9514-9900 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Comment: | Amino acid-binding protein | ||||||||||
| Protein Sequence: | MFDVHVVLDN QIGQLALLGK TLGNKGIGLE GGGIFTVGDE CHAHFLVEQG KEAKIALEQA GLLVLAIRTP LIRKLKQEKP GELGEIARVL AENNINILVQ YSDHANQLIL ITDNDSMAAS VTLPWAIK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | jemC | JemC | Tn10_p | 228 | 9908-10594 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | HTH ArsR-type DNA-binding domain (InterPro:IPR001845) | ||||||||||
| Comment: | N-terminal 100 amino acids similar to bacterial transcriptional repressors and metal-binding proteins (PMID:10781570) | ||||||||||
| Protein Sequence: | MANLIRKEVT FESSIAAIGA AMSDISRVKI LSALMDGRAW TATELSSVAN ISASTASSHL SKLLDCQLIT VVAQGKHRYF RLAGKDIAEL MESMMGISLN HGVHARVSTP VHLRKARTCY DHLAGEVAVK IYDSLCQQQW ITENGSMITL SGIQYFHEMG IDVPSKHSRK ICCACLDWSE RRFHLGGYVG AALFSLYESK GWLTRHLGYR EVTITEKGYA AFKTHFHI |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tetR (ARO:3003479) | TetR | Tn10_p | 208 | 10572-11198 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target alteration (ARO:0001001)||antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Transposase Chemistry: | repressor of the tetracycline resistance element | ||||||||||
| Target: | tetracycline antibiotic (ARO:3000050) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3003479 | ||||||||||
| Protein Sequence: | MMSRLDKSKV INSALELLNE VGIEGLTTRK LAQKLGVEQP TLYWHVKNKR ALLDALAIEM LDRHHTHFCP LEGESWQDFL RNNAKSFRCA LLSHRDGAKV HLGTRPTEKQ YETLENQLAF LCQQGFSLEN ALYALSAVGH FTLGCVLEDQ EHQVAKEERE TPTTDSMPPL LRQAIELFDH QGAEPAFLFG LELIICGLEK QLKCESGS |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tet(B) (ARO:3000166) | Tet(B) | Tn10_p | 401 | 11277-12482 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Transposase Chemistry: | antibiotic efflux (ARO:0010000) | ||||||||||
| Target: | tetracycline antibiotic (ARO:3000050) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3000166 (bitscore:786)||99.9% identical to TnCentral reference sequence for tet(B) (Tn10)||Synonym: tetB | ||||||||||
| Protein Sequence: | MNSSTKIALV ITLLDAMGIG LIMPVLPTLL REFIASEDIA NHFGVLLALY ALMQVIFAPW LGKMSDRFGR RPVLLLSLIG ASLDYLLLAF SSALWMLYLG RLLSGITGAT GAVAASVIAD TTSASQRVKW FGWLGASFGL GLIAGPIIGG FAGEISPHSP FFIAALLNIV AFLVVMFWFR ETKNTRDNTD TEVGVETQSN SVYITLFKTM PILLIIYFSA QLIGQIPATV WVLFTENRFG WNSMMVGFSL AGLGLLHSVF QAFVAGRIAT KWGEKTAVLL GFIADSSAFA FLAFISEGWL VFPVLILLAG GGIALPALQG VMSIQTKSHQ QGALQGLLVS LTNATGVIGP LLFAVIYNHS LPIWDGWIWI IGLAFYCIII LLSMTFMLTP QAQGSKQETS A |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tetC_p | N/A | Tn10_p | 84 | 12595-12849 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Sequence Family: | Bacterial regulatory proteins, tetR family (Pfam:PF00440) | ||||||||||
| Target: | tetracycline antibiotic (ARO:3000050) | ||||||||||
| Comment: | no matchin CARD||identical to C-terminal 84 amino acids of tetC (Tn10) | ||||||||||
| Protein Sequence: | MPIIGWNELE KISLEYITGK VNAIVSKLIQ ENQLKAYDDD VLKNLLNGWF MHIAIHAKNL KELADKKGQF IAIYRGFLLS LKDK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | insAB | InsAB | IS1R | 232 | 12962-13659 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | fusion protein from -1 programmed frameshifting between insA and insB on aaaaaa sequence | ||||||||||
| Protein Sequence: | MASVSISCPS CSATDGVVRN GKSTAGHQRY LCSHCRKTWQ LQFTYTASQP GTHQKIIDMA MNGVGCRATA RIMGVGLNTI FRHLKKLRPQ SVTSRIQPGS DVIVCAEMDE QWGYVGAKSR QRWLFYAYDR LRKTVVAHVF GERTMATLGR LMSLLSPFDV VIWMTDGWPL YESRLKGKLH VISKRYTQRI ERHNLNLRQH LARLGRKSLS FSKSVELHDK VIGHYLNIKH YQ |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | insA | InsA | IS1R | 91 | 12962-13237 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Comment: | regulatory protein | ||||||||||
| Protein Sequence: | VASVSISCPS CSATDGVVRN GKSTAGHQRY LCSHCRKTWQ LQFTYTASQP GTHQKIIDMA MNGVGCRATA RIMGVGLNTI FRHLKNSGRS R |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | insB | InsB | IS1R | 167 | 13156-13659 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | downstream catalytic domain | ||||||||||
| Protein Sequence: | MPGNRPHYGR WPQHDFPPFK KLRPQSVTSR IQPGSDVIVC AEMDEQWGYV GAKSRQRWLF YAYDRLRKTV VAHVFGERTM ATLGRLMSLL SPFDVVIWMT DGWPLYESRL KGKLHVISKR YTQRIERHNL NLRQHLARLG RKSLSFSKSV ELHDKVIGHY LNIKHYQ |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merR | MerR | Tn21.1 | 144 | 13766-14200 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | activator-repressor of mer operon | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MENNLENLTI GVFAKAAGVN VETIRFYQRK GLLREPDKPY GSIRRYGEAD VVRVKFVKSA QRLGFSLDEI AELLRLDDGT HCEEASSLAE HKLKDVREKM ADLARMETVL SELVCACHAR KGNVSCPLIA SLQGEAGLAR SAMP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merT | MerT | Tn21.1 | 116 | 14272-14622 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | cytosolic mercuric ion transport protein | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSEPQNGRGA LFAGGLAAIL ASTCCLGPLV LVALGFSGAW IGNLTVLEPY RPLFIGAALV ALFFAWKRIY RPVQACKPGE VCAIPQVRAT YKLIFWIVAV LVLVALGFPY VVPFFY |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merP | MerP | Tn21.1 | 91 | 14636-14911 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MKKLFASLAL AAAVAPVWAA TQTVTLAVPG MTCAACPITV KKALSKVEGV SKVDVGFEKR EAVVTFDDTK ASVQKLTKAT ADAGYPSSVK Q |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merC | MerC | Tn21.1 | 140 | 14947-15369 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | transmembrane protein mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MGLMTRIADK TGALGSVVSA MGCAACFPAL ASFGAAIGLG FLSQYEGLFI SRLLPLFAAL AFLANALGWF SHRQWLRSLL GMIGPAIVFA ATVWLLGNWW TANLMYVGLA LMIGVSIWDF VSPAHRRCGP DGCELPAKRL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merA | MerA | Tn21.1 | 564 | 15421-17115 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercuric ion reductase | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSTLKITGMT CDSCAVHVKD ALEKVPGVQS ADVSYAKGSA KLAIEVGTSP DALTAAVAGL GYRATLADAP SVSTPGGLLD KMRDLLGRND KTGSSGALHI AVIGSGGAAM AAALKAVEQG ARVTLIERGT IGGTCVNVGC VPSKIMIRAA HIAHLRRESP FDGGIAATTP TIQRTALLAQ QQARVDELRH AKYEGILEGN PAITVLHGSA RFKDNRNLIV QLNDGGERVV AFDRCLIATG ASPAVPPIPG LKDTPYWTST EALVSETIPK RLAVIGSSVV ALELAQAFAR LGAKVTILAR STLFFREDPA IGEAVTAAFR MEGIEVREHT QASQVAYING EGDGEFVLTT AHGELRADKL LVATGRAPNT RKLALDATGV TLTPQGAIVI DPGMRTSVEH IYAAGDCTDQ PQFVYVAAAA GTRAAINMTG GDAALNLTAM PAVVFTDPQV ATVGYSEAEA HHDGIKTDSR TLTLDNVPRA LANFDTRGFI KLVVEEGSGR LIGVQAVAPE AGELIQTAAL AIRNRMTVQE LADQLFPYLT MVEGLKLAAQ TFNKDVKQLS CCAG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merD | MerD | Tn21.1 | 120 | 17133-17495 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | secondary regulatory protein | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSAYTVSQLA HNAGVSVHIV RDYLVRGLLR PVACTTGGYG VFDDAALQRL CFVRAAFEAG IGLDALARLC RALDAADGAQ AAAQLAVLRQ LVERRRAALA HLDAQLASMP AERAHEEALP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merE | MerE | Tn21.1 | 78 | 17492-17728 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Comment: | similar to urf-1 in pKLH2 (GenBank AF213017), pKLH272 (Genbank Y08992), pMER610 (GenBank Y08993), pKLH210 (GenBank Y10102), Tn5036 (Genbank Y09025), orf1 in Tn501 (GenBank Z00027), and urf-1 in Tn5041 (GenBank X98999) | ||||||||||
| Protein Sequence: | MNAPDKLPPE TRQPVSGYLW GALAVLTCPC HLPILAAVLA GTTAGAFLGE HWGVAALALT GLFVLAVTRL LRAFRGGS |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | urfM 5'-end | N/A | Tn21.2 | 223 | 17725-18395 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Comment: | urfMORF interrupted by insertion of In2 | ||||||||||
| Protein Sequence: | MTSSQPAGWT AAELAQAAAR GQLDLHYQPL VDLRDHRIAG AEALMRWRHP RLGLLPPGQF LPLAESFGLM PEIGAWVLGE ACRQMHKWQG PAWQPFRLAI NVSASQVGPT FDDEVKRVLA DMALPAELLE IELTESVAFG NPALFASFDA LRAIGVRFAA DDFGTGYSCL QHLKCCPITT LKIDQSFVAR LPDDARDQTI VRAVIQLAHG LGMDVIFRRR LHQ |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA_p | N/A | In_Tn21.1 | 434 | 18471-19775 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | integrase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | Contains the first 429 amino acids of tniA (In2)||probably truncated by insertion of IS26 | ||||||||||
| Protein Sequence: | MLNTRVHQSE VSMATDTPRI PEQGVATLPD EAWERARRRA EIISPLAQSE TVGHEAADMA AQALGLSRRQ VYVLIRRARQ GSGLVTDLVP GQSGGGKGKG RLPEPVERVI HELLQKRFLT KQKRSLAAFH REVTQVCKAQ KLRVPARNTV ALRIASLDPR KVIRRREGQD AARDLQGVGG EPPAVTAPLE QVQIDHTVID LIVVDDRDRQ PIGRPYLTLA IDVFTRCVLG MVVTLEAPSA VSVGLCLVHV ACDKRPWLEG LNVEMDWQMS GKPLLLYLDN AAEFKSEALR RGCEQHGIRL DYRPLGQPHY GGIVERIIGT AMQMIHDELP GTTFSNPDQR GDYDSENKAA LTLRELERWL TLAVGTYHGS VHNGLLQPPA ARWAEAVARV GVPAVVTRAT SFLVDFLPIL RRTLTRTGFV IDHIHYYADG HCCK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 19822-20526 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | SDR family oxidoreductase | SDR family oxidoreductase | In_Tn21.1 | 287 | 20984-21847 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | WP_000612791.1 | ||||||||||
| Protein Sequence: | MIPNSENKRV WFITGASKGL GYAFTCAALK AGDKVVAVAR TIDNLAKLEE TYQESLLPLN LDVTDREAVF STVETAVKHF GRLDIVVNNA GIMTMGMIEE LNESDARKLM DTNFFGALWV CQAVMPYLRS QRSGHIIQIT SIGAIISGPM SGIYSASKFA LEGMSEALAK EAEHFGVKLT MVEPGGYWTD LYTSMSYSNP LDSYGTLRDE LAKQYSEDSV DSDPSLAAEA LMKLVASNNP PLRLILGSMV YDLAMDTLKA RMATWEEWEA VSRASEKAIP APERYGV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | GrpB domain protein | GrpB domain protein | In_Tn21.1 | 81 | 21885-22130 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | GrpB (Pfam:PF04229) | ||||||||||
| Protein Sequence: | MKIEIMEYNP DWTKNFEEEK IKLLHFFGSH AVAIEHIGST AIPNQRAKPV IDIFIGVSPF AELPFISAFL MQRSITTLRQ I |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | sul3 (ARO:3000413) | Sul3 | In_Tn21.1 | 263 | 22599-23390 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | sulfonamide resistant sul (ARO:3004238) | ||||||||||
| Target: | sulfone antibiotic (ARO:3003401)||sulfonamide antibiotic (ARO:3000282) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000413 | ||||||||||
| Protein Sequence: | MSKIFGIVNI TTDSFSDGGL YLDTDKAIEH ALHLVEDGAD VIDLGAASSN PDTTEVGVVE EIKRLKPVIK ALKEKGISIS VDTFKPEVQS FCIEQKVDFI NDIQGFPYPE IYSGLAKSDC KLVLMHSVQR IGAATKVETN PEEVFTSMME FFKERIAALV EAGVKRERII LDPGMGFFLG SNPETSILVL KRFPEIQEAF NLQVMIAVSR KSFLGKITGT DVKSRLAPTL AAEMYAYKKG ADYLRTHDVK SLSDALKISK ALG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnp IS256 family | Tnp IS256 family | In_Tn21.1 | 202 | 23715-24323 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | tranposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MEFYPSCIEK GMRSERALKL AIAEMYVKGV STRRVSDIVE ILCGTEVSSS QVSRLAKELD EEITSWKAQP VGQIQYLVLD ATYESVRVGS HVVKQALLVA IGVDYSGNRH ILDAEVANSE AEVNWRSFLE GLVRRGMHGL RMITSDDHSG LRAAIDAVFP GILWQRCQFH LQQNAHSYVT KKDEIPLIAA DIRKVFNRNM SR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacL (ARO:3005098) | QacL | In_Tn21.1 | 110 | 24570-24902 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | small multidrug resistance (SMR) antibiotic efflux pump (ARO:0010003) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | subunit of the qac multidrug efflux pump||strict match to reference sequence for ARO:3005098 (bitscore: 202) | ||||||||||
| Protein Sequence: | MKNWLFLAIA IFGEVVATSA LKSSHGFTKL VPSVVVVAGY GLAFYFLSLA LKSIPVGIAY AVWAGLGIVL VAAIAWIFHG QKLDLWAFVG MGLIVSGVAV LNLLSKVSAH |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aadA (ARO:3002601) | AadA | In_Tn21.1 | 263 | 25072-25863 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'') (ARO:3004275) | ||||||||||
| Transposase Chemistry: | aminoglycoside nucleotidyltransferase | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002601||Synonyms: aadA1-pm aadA, aadA1, aad(3'')(9) | ||||||||||
| Protein Sequence: | MREAVIAEVS TQLSEVVGVI ERHLEPTLLA VHLYGSAVDG GLKPHSDIDL LVTVTVRLDE TTRRALINDL LETSASPGES EILRAVEVTI VVHDDIIPWR YPAKRELQFG EWQRNDILAG IFEPATIDID LAILLTKARE HSVALVGPAA EELFDPVPEQ DLFEALNETL TLWNSPPDWA GDERNVVLTL SRIWYSAVTG KIAPKDVAAD WAMERLPAQY QPVILEARQA YLGQEEDRLA SRADQLEEFV HYVKGEITKV VGK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | cmlA6 (ARO:3002696) | CmlA6 | In_Tn21.1 | 419 | 25956-27215 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | phenicol antibiotic (ARO:3000387) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002696 (bitscore: 819) | ||||||||||
| Protein Sequence: | MSSKNFSWRY SLAATVLLLS PFDLLASLGM DMYLPAVPFM PNALGTTAST IQLTLTTYLV MIGAGQLLFG PLSDRLGRRP VLLGGGLAYV VASMGLALTS SAEVFLGLRI LQACGASACL VSTFATVRDI YAGREESNVI YGILGSMLAM VPAVGPLLGA LVDMWLGWRA IFAFLGLGMI AASAAAWRFW PETRVQRVAG LQWSQLLLPV KCLNFWLYTL CYAAGMGSFF VFFSIAPGLM MGRQGVSQLG FSLLFATVAI AMVFTARFMG RVIPKWGSPS VLRMGMGCLI AGAVLLAITE IWALQSVLGF IAPMWLVGIG VATAVSVAPN GALRGFDHVA GTVTAVYFCL GGVLLGSIGT LIISLLPRNT AWPVVVYCLT LATVVLGLSC VSRVKGSRGQ GEHDVVALQS AESTSNPNR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aadA2 (ARO:3002602) | AadA2 | In_Tn21.1 | 259 | 27477-28256 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'') (ARO:3004275) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002602 (bitscore: 520) | ||||||||||
| Protein Sequence: | VTIEISNQLS EVLSVIERHL ESTLLAVHLY GSAVDGGLKP YSDIDLLVTV AVKLDETTRR ALLNDLMEAS AFPGESETLR AIEVTLVVHD DIIPWRYPAK RELQFGEWQR NDILAGIFEP AMIDIDLAIL LTKAREHSVA LVGPAAEEFF DPVPEQDLFE ALRETLKLWN SQPDWAGDER NVVLTLSRIW YSAITGKIAP KDVAADWAIK RLPAQYQPVL LEAKQAYLGQ KEDHLASRAD HLEEFIRFVK GEIIKSVGK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | DUF1010 family protein | DUF1010 family protein | In_Tn21.1 | 96 | 28274-28564 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | DUF1010 (Pfam:PF06231) | ||||||||||
| Protein Sequence: | MFIQTAFSFS GVIQCLFCLF SGLRLHGLRR FSVFLASSPC VASASSYRFC SAVPPRWRSV FSRLAPVAKF KLSVLASGSN ISVKPTRILR SAYLAR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | dfrA12 (ARO:3002858) | DfrA12 | In_Tn21.1 | 165 | 28676-29173 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | trimethoprim resistant dihydrofolate reductase dfr (ARO:3001218) | ||||||||||
| Target: | diaminopyrimidine antibiotic (ARO:3000171) | ||||||||||
| Comment: | 100% identity with reference sequence for ARO:3002858 (bitscore: 339)||Synonyms: | ||||||||||
| Protein Sequence: | MNSESVRIYL VAAMGANRVI GNGPNIPWKI PGEQKIFRRL TEGKVVVMGR KTFESIGKPL PNRHTLVISR QANYRATGCV VVSTLSHAIA LASELGNELY VAGGAEIYTL ALPHAHGVFL SEVHQTFEGD AFFPMLNETE FELVSTETIQ AVIPYTHSVY ARRNG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI1 | IntI1 | In_Tn21.1 | 337 | 29318-30331 | + |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 1 | ||||||||||
| Function: | Integrase | ||||||||||
| Sequence Family: | Class 1 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MKTATAPLPP LRSVKVLDQL RERIRYLHYS LPTEQAYVHW VRAFIRFHGV RHPATLGSSE VEAFLSWLAN ERKVSVSTHR QALAALLFFY GKVLCTDLPW LQEIGRPRPS RRLPVVLTPD EVVRILGFLE GEHRLFAQLL YGTGMRISEG LQLRVKDLDF DHGTIIVREG KGSKDRALML PESLAPSLRE QLSRARAWWL KDQAEGRSGV ALPDALERKY PRAGHSWPWF WVFAQHTHST DPRSGVVRRH HMYDQTFQRA FKRAVEQAGI TKPATPHTLR HSFATALLRS GYDIRTVQDL LGHSDVSTTM IYTHVLKVGG AGVRSPLDAL PPLTSER |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpM | TnpM | Tn21.1 | 116 | 30534-30884 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Inhibitor | ||||||||||
| Function: | transposition regulator||reported to enhance Tn21 transposition and suppress resolution of cointegrate replicons in vivo | ||||||||||
| Comment: | 3'-end of urfM ORF, which is interrupted by insertion of In2||inhibits tranposition probably by inhibiting resolution | ||||||||||
| Protein Sequence: | MEVVAEGVET PDCLAWLRQA GCDTVQGFLF ARPMPAAAFV GFVNQWRNTT MNANEPSTSC CVCCKEIPLD AAFTPEGAEY VEHFCGLECY QRFQARASTA TETSVKPDAC DSPPSG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR 5'-end | N/A | Tn21.1 | 7 | 31010-31032 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Comment: | tnpR ORF interrupted by IS26 insertion | ||||||||||
| Protein Sequence: | MTGQRIG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 31104-31808 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR 3'-end | N/A | Tn21.1 | 179 | 31861-32398 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Comment: | tnpR ORF interrupted by IS26 insertion | ||||||||||
| Protein Sequence: | YQGQHLRPEP GTATGRRQG* SRF*RQGIRQ GCQASATGSA DKLRPHRRHR GGA*HGSPGA QSR*FAPDRA NADTTRRAYR IRQGTPQFYW RRLSDGEPDA LGDGRVRRVR ARPDPRASAR GYCARQATRG LPWQEEIPVV *AYCRTAPTC RGWRAKDQAC S*IRNQSRNP VSILENGSV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn21.1 | 988 | 32401-35367 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposition, DNA-mediated (GO:0006313) | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | identical to TnAs3 tnpA | ||||||||||
| Protein Sequence: | MPRRSILSAA ERESLLALPD SKDDLIRHYT FNDTDLSIIR QRRGPANRLG FAVQLCYLRF PGVILGVDEL PFPPLLKLVA DQLKVGVESW NEYGQREQTR REHLSELQTV FGFRPFTMSH YRQAVQMLTE LAMQTDKGIV LASALIGHLR RQSVILPALN AVERASAEAI TRANRRIYDA LAEPLADAHR RRLDDLLKRR DNGKTTWLAW LRQSPAKPNS RHMLEHIERL KAWQALDLPT GIERLVHQNR LLKIAREGGQ MTPADLAKFE PQRRYATLVA LATEGMATVT DEIIDLHDRI LGKLFNAAKN KHQQQFQASG KAINAKVRLY GRIGQALIDA KQSGRDAFAA IEAVMSWDSF AESVTEAQKL AQPDDFDFLH RIGESYATLR RYAPEFLAVL KLRAAPAAKN VLDAIEVLRG MNTDNARKLP ADAPTGFIKP RWQKLVMTDA GIDRRYYELC ALSELKNSLR SGDIWVQGSR QFKDFEDYLV PPEKFTSLKQ SSELPLAVAT DCEQYLHERL TLLEAQLATV NRMAAANDLP DAIITESGLK ITPLDAAVPD TAQALIDQTA MVLPHVKITE LLLEVDEWTG FTRHFTHLKS GDLAKDKNLL LTTILADAIN LGLTKMAESC PGTTYAKLAW LQAWHTRDET YSTALAELVN AQFRHPFAGH WGDGTTSSSD GQNFRTASKA KSTGHINPKY GSSPGRTFYT HISDQYAPFH TKVVNVGLRD STYVLDGLLY HESDLRIEEH YTDTAGFTDH VFALMHLLGF RFAPRIRDLG DTKLYIPKGD AAYDALKPMI GGTLNIKHVR AHWDEILRLA TSIKQGTVTA SLMLRKLGSY PRQNGLAVAL RELGRIERTL FILDWLQSVE LRRRVHAGLN KGEARNALAR AVFFNRLGEI RDRSFEQQRY RASGLNLVTA AIVLWNTVYL ERAAHALRGN GHAVDDSLLQ YLSPLGWEHI NLTGDYLWRS SAKIGAGKFR PLRPLQPA |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| IS26-MH257753 | IS26 | insertion sequence | 6027-6846 | + | 820 |
| Tn10_p-MH626558 | Tn10_p | transposon | 6842-12907 | + | 6066 |
| IS10_p-MH626558 | IS10_p | insertion sequence | 6847-7199 | + | 353 |
| IS1R-J01730 | IS1R | insertion sequence | 12907-13674 | + | 768 |
| Tn21.1-MH257753 | Tn21.1 | transposon | 13733-35400 | + | 21668 |
| In_Tn21.1-MH257753 | In_Tn21.1 | integron | 18366-30533 | + | 12168 |
| IS26-MH257753 | IS26 | insertion sequence | 19759-20578 | + | 820 |
| IS26-MH257753 | IS26 | insertion sequence | 31041-31860 | + | 820 |
| Name | Associated TE | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|---|
| IRL | IS26 | 6027-6040 | + | 14 | GGCACTGTTG CAAA |
| IRR | IS26 | 6833-6846 | - | 14 | GGCACTGTTG CAAA |
| IRR | IS10_p | 7178-7199 | - | 22 | CTGAGAGATC CCCTCATAAT TT |
| IRL | IS1R | 12907-12929 | + | 23 | GGTGATGCTG CCAACTTACT GAT |
| IRR | IS1R | 13652-13674 | - | 23 | GGTAATGACT CCAACTTATT GAT |
| IRL | Tn21.1 | 13733-13770 | + | 38 | GGGGGCACCT CAGAAAACGG AAAATAAAGC ACGCTAAG |
| IRt | In_Tn21.1 | 18366-18398 | + | 33 | TGTCATTTTC AGAAGACGAC TGCACCAGTT GAT |
| repeat t1 | In_Tn21.1 | 18374-18392 | + | 19 | TCAGAAGACG ACTGCACCA |
| repeat t2 | In_Tn21.1 | 18414-18432 | - | 19 | TCAGGAGCTG GCTGCACAA |
| repeat t3 | In_Tn21.1 | 18443-18462 | + | 20 | TCAGAAGTGA TCTGCACCAA |
| repeat t4 | In_Tn21.1 | 18475-18493 | + | 19 | TCAATACTCG TGTGCACCA |
| IRL | IS26 | 19759-19772 | + | 14 | GGCACTGTTG CAAA |
| IRR | IS26 | 20565-20578 | - | 14 | GGCACTGTTG CAAA |
| repeat i4 | In_Tn21.1 | 30414-30432 | - | 19 | TCAGCGGACG CAGGGAGGA |
| repeat i3 | In_Tn21.1 | 30442-30460 | - | 19 | TCAGGCAACG ACGGGCTGC |
| repeat i2 | In_Tn21.1 | 30484-30502 | - | 19 | TCAGAAGCCG ACTGCACTA |
| IRi | In_Tn21.1 | 30501-30533 | - | 33 | TGTCGTTTTC AGAAGACGGC TGCACTGAAC GTC |
| repeat i1 | In_Tn21.1 | 30507-30525 | - | 19 | TCAGAAGACG GCTGCACTG |
| IRL | IS26 | 31041-31054 | + | 14 | GGCACTGTTG CAAA |
| IRR | IS26 | 31847-31860 | - | 14 | GGCACTGTTG CAAA |