Mobile element type:
Transposon
Name:
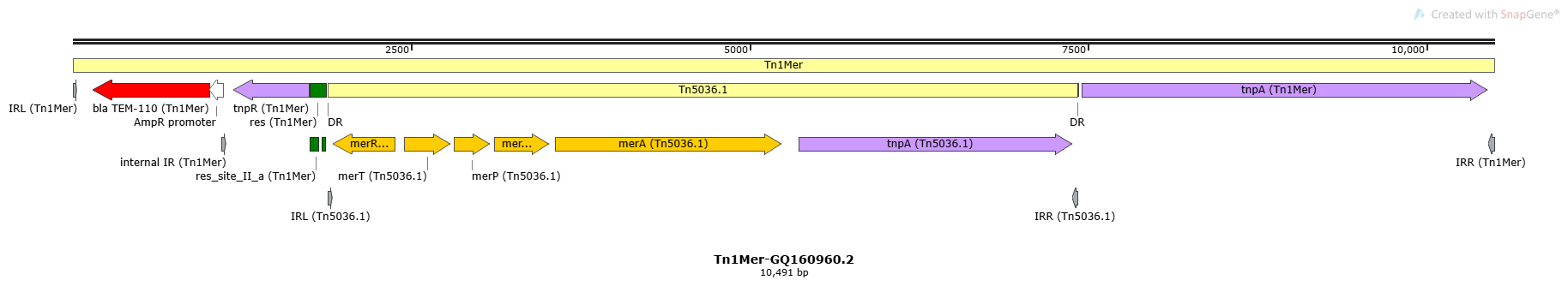
Tn1Mer
Synonyms:
Tn7238
Accession:
Tn1Mer-GQ160960.2
Family:
Tn3
Group:
Tn3
First isolate:
ND
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Serratia marcescens
Date of Isolation:
None
Country:
France
Molecular Source:
plasmid R934
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRL | 1-38 | + | 38 | GGGGTCTGAC GCTCAGTGGA ACGAAAACTC ACGTTAAG |
| internal IR | 1103-1140 | + | 38 | GGGGTTCCGC GCACATTTCC CCGAAAAGTG CCACCTGA |
| IRR | 10454-10491 | - | 38 | GGGGTCTGAC GCTCAGTGGA ACGAAAACTC ACGTTAAG |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| res | Tn1Mer | 1750-1870 | + | 121 | AAATGTACCT TAAATCGAAT ATCAGACACG ATGTGTCTAT TATGCCAAAA TGACGATTTA ATGGACACTC AAACGAAGCC GTTTTACTAT GTCTGATAAT TTATAACATT TCGGACGGTT G |
| res_site_III_a | Tn1Mer | 1753-1775 | + | 23 | TGTACCTTAA ATCGAATATC AGA |
| res_site_II_a | Tn1Mer | 1781-1816 | + | 36 | TGTGTCTATT ATGCCAAAAT GACGATTTAA TGGACA |
| res_site_I | Tn1Mer | 1839-1867 | + | 29 | TGTCTGATAA TTTATAACAT TTCGGACGG |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| bla TEM-110 (ARO:3000973) | Tn1Mer | 148-1008 | - | Passenger Gene | Antibiotic Resistance |
| tnpR | Tn1Mer | 1191-1748 | - | Accessory Gene | Resolvase |
| merR | Tn5036.1 | 1921-2376 | - | Passenger Gene | Heavy Metal Resistance |
| merT | Tn5036.1 | 2448-2798 | + | Passenger Gene | Heavy Metal Resistance |
| merP | Tn5036.1 | 2814-3089 | + | Passenger Gene | Heavy Metal Resistance |
| merC | Tn5036.1 | 3117-3524 | + | Passenger Gene | Heavy Metal Resistance |
| merA | Tn5036.1 | 3563-5245 | + | Passenger Gene | Heavy Metal Resistance |
| tnpA | Tn5036.1 | 5364-7391 | + | Transposase | |
| tnpA | Tn1Mer | 7453-10458 | + | Transposase |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | bla TEM-110 (ARO:3000973) | Bla TEM-110 | Tn1Mer | 286 | 148-1008 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | TEM beta-lactamase (ARO:3000014) | ||||||||||
| Target: | cephalosporin (ARO:0000032)||penem (ARO:3003706)||monobactam (ARO:0000004)||penam (ARO:3000008) | ||||||||||
| Comment: | hydrolyzes the beta-lactam bond in penicillins and cephalosporins||perfectmatch to reference sequence for ARO:300973 | ||||||||||
| Protein Sequence: | MSIQHFRVAL IPFFAAFCFP VFAHPETLVK VKDAEDQLGA RVGYIELDLN SGKILESFRP EERFPMMSTF KVLLCGAVLS RVDAGQEQLG RRIHYSQNDL VEYSPVTEKH LTDGMTVREL CSAAITMSDN TAANLLLTTI GGPKELTAFL HNMGDHVTRL DRWEPELNEA IPNDERDTTM PAAMATTLRK LLTGELLTLA SRQQLIDWME ADKVAGPLLR SALPAGWFIA DKSGAGERGS RGIIAALGPD GKPSRIVVIY MTGSQATMDE RNRQIAEIGA SLIKHW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | Tn1Mer | 185 | 1191-1748 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Function: | recombinase acitivity (GO:0000150) | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MRIFGYARVS TSQQSLDIQI RALKDAGVKA NRIFTDKASG SSTDREGLDL LRMKVEEGDV ILVKKLDRLG RDTADMIQLI KEFDAQGVAV RFIDDGISTD GDMGQMVVTI LSAVAQAERR RILERTNEGR QEAKLKGIKF GRRRTVDRNV VLTLHQKGTG ATEIAHQLSI ARSTVYKILE DERAS |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merR | MerR | Tn5036.1 | 151 | 1921-2376 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | activator-repressor of mer operon | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MQINFENLTI GVFAKAAGVN VETIRFYQRK GLLPEPDKPY GSIRRYGEAD VTRVRFVKSA QRLGFSLDEI AELLRLEDGT HCEEASGLAE HKLKDVREKM ADLARMEAVL SELVCACHAR KGNVSCPLIA SLQDGTKLAA SARGSHGVTT P |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merT | MerT | Tn5036.1 | 116 | 2448-2798 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercuric ion transport protein | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MSEPQNGRGA LFAGGLAAIL ASACCLGPLV LIALGFSGAW IGNLTVLEPY RPIFIGAALV ALFFAWRRIY RPAQACKPGE VCAIPQVRAT YKLIFWIVAV LVLVALGFPY VMPFFY |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merP | MerP | Tn5036.1 | 91 | 2814-3089 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercury transport | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MKKLFAALAL AAVVAPVWAT TQTVTLSVPG MTCSTCPITV KKAISKVEGV SKIDVTFETR EAVVTFDDAK TSVQKLTKAT GDAGYPSSVK Q |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merC | MerC | Tn5036.1 | 135 | 3117-3524 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | mercurry ion transport Go:0015694 | ||||||||||
| Target: | Mercury | ||||||||||
| Protein Sequence: | MGLITRIADK AGALGSVVSA MGCAACFPAI ASLGAAIGLG FLQEYEGLFI SKLLPLFAVV ALLANALGWL SHRQWYRSLL GMVGPAIVLA AMLLFFGNWW TANLLYVGLA LMIGVSIWDL LSPANRRCEL PPKHG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merA | MerA | Tn5036.1 | 560 | 3563-5245 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Function: | response to mercury ion (GO:0046689) | ||||||||||
| Target: | Mercury | ||||||||||
| Comment: | Hg (II) mercuric ion reductase||Synonym= | ||||||||||
| Protein Sequence: | MYLNITGMTC DSCATHVKDA LEKVPGVLSA LVSYPKGSAQ LATDPGTSPE ALTAAVAGLG YKATPADAPS TSARGRLLGK ALGWLGGGDK AGGDGDGLHV AVIGSGGAAM AAALKAVEQG ANVTLIERGT IGGTCVNVGC VPSKIMIRAA HIAHLRRESP FDSGIAATVP AIDRSKLLAQ QQARVDELRH AKYEGILDSN PAITVLHGEA RFKDDQSLAV RLNDGGERVV AFDRCLVATG ASPAVPPIPG LKESPYWTST EALVSDTIPE RLAVIGSSVV ALELAQAFAR LGSKVTILAR STLFFREDPA IGEAVTAAFR AEGIKVLEHT QASQVAHVDG EFVLTTGYGE IRADQLLVAT GRAPNTRSLA LEAAGVAANA QGAIVIDKGM RTSTPHIYAA GDCTDQPQFV YVAAAAGTRA AINMTGGDAA LDLTAMPAVV FTDPQVATVG YSEAEAHHDG IETDSRTLTL DNVPRALANF DTRGFIKLVI EEGSGRLIGV QAVAPEAGEL IQTAVLAIRN RMTVQELADQ LFPYLTMVEG LKLAAQTFSK DVKQLSCCAG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn5036.1 | 675 | 5364-7391 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase activity (GO:0004803) | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | internal deletion | ||||||||||
| Protein Sequence: | MTLADLAKFE VQRRYATLVA LAIEGMATVT DEIIDLHDRI IGKLFNAAKN KHQQQFQASG KAINDKVRMY GRIGQALIEA KQSGSDPFAA IEAVMPWDTF AASVTEAQTL ARPADFDFLH HIGESYATLR RYAPQFLGVL KLRAVLDAID MLRGMNSDSA CKVPADAPTA FIKPRWAKLV LTDDGIDRRY YELCALSELK NALRSGDVWV QGSRQFKDFD EYLVPVEKFA TLKLASELPL DAAVPDAAQA MIDQTAMLLP HLKITELLME VDEWTGFTRH FTHLKTSDTA KDKTLLLTTI LADAINLGLT KMAESCPGTT YAKLSWLQAW HIRDETYSTA LAELVNAQFR QPFAGNWDDG TTSSSDGQNF RTGSKAESTG HINPKYGSSP GRTFYTHISD QYAPFSAKVV NVGIRDSTYV LDGLLYHESD LRIEEHYTDT AGFTDHVFGL MHLLGFRFAP RIRDLGETKL FIPKGDAAYD ALKPMISSDR LNIKQIRAHW DEILRLATSI KQGTVTASLM LRKLGSYPRQ NGLAVALREL GRIEHTLFIL DWLQSVELRR RVHAGLNKGE ARNALARAVF FYRLGEIRDR SFEQQRYRAS GLNLVTAAIV LWNTVYLERA TSALRGNGTA LDDTLLQYLS PLGWEHINLT GDYLWRSSAK VGAGKFRPLR PLPPA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn1Mer | 1001 | 7453-10458 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase activity (GO:0004803) | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MPVDFLTTEQ TESYGRFTGE PDELQLARYF HLDEADKEFI GKSRGDHNRL GIALQIGCVR FLGTFLTDMN HIPSGVRHFT ARQLGIRDIT VLAEYGQREN TRREHAALIR QHYQYREFAW PWTFRLTRLL YTRSWISNER PGLLFDLATG WLMQHRIILP GATTLTRLIS EVREKATLRL WNKLALIPSA EQRSQLEMLL GPTDCSRLSL LESLKKGPVT ISGPAFNEAI ERWKTLNDFG LHADNLSTLP AVRLKNLARY AGMTSVFNIA RMSPQKRMAV LVAFVLAWET LALDDALDVL DAMLAVIIRD ARKIGQKKRL RSLKDLDKSA LALASACSYL LKEETPDESI RAEVFSYIPR QKLAEIITLV REIARPSDDN FHEEMVEQYG RVRRFLPHLL NTVKFSSAPA GVTTLNACDY LSREFSSRRQ FFDDAPTEII SRSWKRLVIN KEKHITRRGY TLCFLSKLQD SLRRRDVYVT GSNRWGDPRA RLLQGADWQA NRIKVYRSLG HPTDPQEAIK SLGHQLDSRY RQVAARLGEN EAVELDVSGP KPRLTISPLA SLDEPDSLKR LSKMISDLLP PVDLTELLLE INAHTGFADE FFHASEASAR VDDLPVSISA VLMAEACNIG LEPLIRSNVP ALTRHRLNWT KANYLRAETI TSANARLVDF QATLPLAQIW GGGEVASADG MRFVTPVRTI NAGPNRKYFG NNRGITWYNF VSDQYSGFHG IVIPGTLRDS IFVLEGLLEQ ETGLNPTEIM TDTAGTSELV FGLFWLLGYQ FSPRLADAGA SVFWRMGHDA NYGVLNDIAR GQSDPRKIVL QWDEMIRTAG SLKLGKVQAS VLVRSLLKSE RPSGLTQAII EVGRINKTLY LLNYIDDEDY RRRILTQLNR GESRHAVARA ICHGQKGEIR KRYTDGQEDQ LGALGLVTNA VVLWNTMYMQ AALDHLRAQG ETLNDEDIAR LSPLCHGHIN MLGHYSFTLA ELVTKGHLRP LKEASEVENV A |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| Tn5036.1-GQ160960.2 | Tn5036.1 | transposon | 1888-7424 | + | 5537 |
| Name | Associated TE | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|---|
| IRL | Tn5036.1 | 1888-1925 | + | 38 | GGGGTCGTCT CAGAATTCGG AAAATAAAGC ACGCTAAG |
| IRR | Tn5036.1 | 7387-7424 | - | 38 | GGGGTCGTCT CAGAAAACGG AAAATAAAGC ACGCTAAG |
1. Pinyon JL, Hall RM.
Evolution of IncP-1alpha plasmids by acquisition of antibiotic and mercuric ion resistance transposons.. Microb Drug Resist. 2011 Sep;17(3):339-43. doi: 10.1089/mdr.2010.0196. Epub 2011 Apr 8. PubMed ID:
21476866
.