Mobile element type:
Transposon
Name:
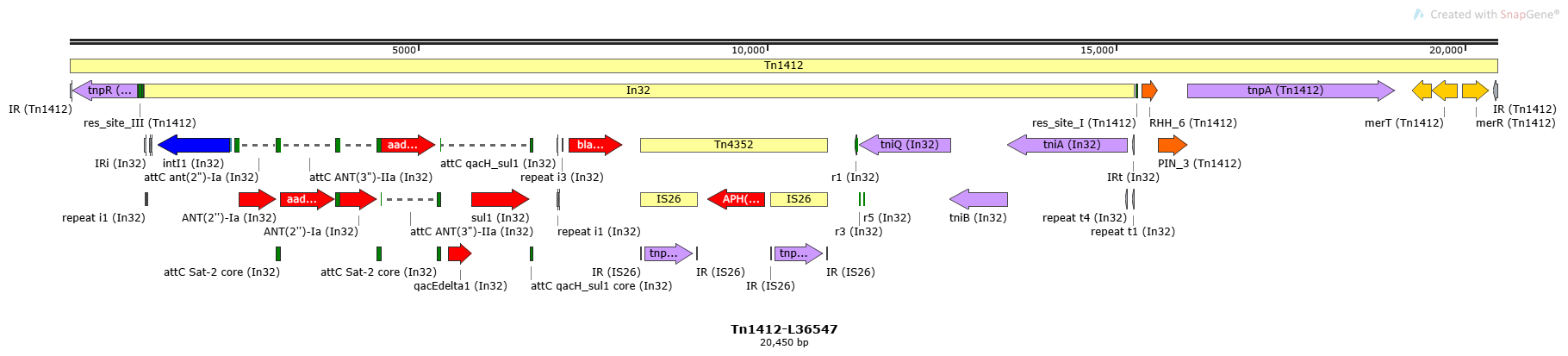
Tn1412
Synonyms:
None
Accession:
Tn1412-L36547
Family:
Tn3
Group:
Tn3
First isolate:
ND
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Pseudomonas aeruginosa 2293E
Date of Isolation:
2001
Country:
None
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IR | 1-39 | + | 39 | GGGGTTTGGG GAGCAATGGA ACCAAAAACC AACGTAAGC |
| IR | 20412-20450 | - | 39 | GGGGTTTGGG GAGCAATGGA ACCAAAAACC AACGTAAGC |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| res_site_III | Tn1412 | 985-1026 | + | 42 | ACTCGACTAC GTTAGTAATA GTTGAACTTT GATTAAGCGT AC |
| res_site_II | Tn1412 | 1029-1059 | + | 31 | GTTATTTGAA CCGTAGCGCG GGGAGCTTAA C |
| attI | In32 | 2366-2421 | + | 56 | TGATGTTATG GAGCAGCAAC GATGTTACGC AGCAGGGCAG TCGCCCTAAA ACAAAG |
| attC ant(2"")-Ia | In32 | 2422-2427 , 2959-3012 | + | 60 | TTAGGCGCCT AACAATTCGT CCAAGCCGAC GCCGCTTCGC GGCGCGGCTT AACTCAGGTG |
| attC Sat-2 core | In32 | 2959-3018 | + | 60 | GCCTAACAAT TCGTCCAAGC CGACGCCGCT TCGCGGCGCG GCTTAACTCA GGTGTTAAAC |
| attC ANT(3"")-IIa | In32 | 3013-3018 , 3815-3868 | + | 60 | TTAAACGTCT AACAATTCGT TCAAGCCGAC GCCGCTTCGC GGCGCGGCTT ACCTTGGCCG |
| attC ANT(3'')-IIa core | In32 | 3815-3868 | + | 54 | GTCTAACAAT TCGTTCAAGC CGACGCCGCT TCGCGGCGCG GCTTACCTTG GCCG |
| attC ant(2"")-Ia | In32 | 3869-3874 , 4406-4459 | + | 60 | TTAGGCGCCT AACAATTCGT CCAAGCCGAC GCCGCTTCGC GGCGCGGCTT AACTCAGGTG |
| attC Sat-2 core | In32 | 4406-4465 | + | 60 | GCCTAACAAT TCGTCCAAGC CGACGCCGCT TCGCGGCGCG GCTTAACTCA GGTGTTAAAC |
| attC ANT(3"")-IIa | In32 | 4460-4465 , 5262-5315 | + | 60 | TTAAACGTCT AACAATTCGT TCAAGCCGAC GCCGCTTCGC GGCGCGGCTT ACCTTGGCCG |
| attC ANT(3'')-IIa core | In32 | 5262-5315 | + | 54 | GTCTAACAAT TCGTTCAAGC CGACGCCGCT TCGCGGCGCG GCTTACCTTG GCCG |
| attC qacH_sul1 | In32 | 5315-5321 , 6599-6632 | + | 41 | GTTAGATGCC TAGCATTCAC CTTCCGGCCG CCCGCTAGCG G |
| attC qacH_sul1 core | In32 | 6599-6632 | + | 34 | GCCTAGCATT CACCTTCCGG CCGCCCGCTA GCGG |
| r1 | In32 | 11263-11276 | - | 14 | CGTATACGTG ACGG |
| r2 | In32 | 11279-11292 | + | 14 | CGATTATGTG ACAG |
| r3 | In32 | 11323-11336 | - | 14 | GGGTTAAGTG ACAA |
| r5 | In32 | 11379-11392 | - | 14 | AATTTCTGTC ACAT |
| res_site_I | Tn1412 | 15281-15308 | + | 28 | GTTCGGCAAA ACCTTCGTTT TGTCGAAC |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tnpR | Tn1412 | 41-970 | - | Accessory Gene | Resolvase |
| intI1 | In32 | 1272-2285 | - | Integron Integrase | Class 1 |
| ANT(2'')-Ia (ARO:3000230) | In32 | 2431-2964 | + | Passenger Gene | Antibiotic Resistance |
| aadA (ARO:3002601) | In32 | 3022-3813 | + | Passenger Gene | Antibiotic Resistance |
| ANT(2'')-Ia (ARO:3000230) | In32 | 3878-4411 | + | Passenger Gene | Antibiotic Resistance |
| aadA (ARO:3002601) | In32 | 4469-5260 | + | Passenger Gene | Antibiotic Resistance |
| qacEdelta1 (ARO:3005010) | In32 | 5424-5771 | + | Passenger Gene | Antibiotic Resistance |
| sul1 (ARO:3000410) | In32 | 5765-6604 | + | Passenger Gene | Antibiotic Resistance |
| bla LCR-1 (ARO:3002997) | In32 | 7158-7940 | + | Passenger Gene | Antibiotic Resistance |
| tnpA | IS26 | 8246-8950 | + | Transposase | |
| APH(3')-Ia (ARO:3002641) | Tn4352 | 9140-9955 | - | Passenger Gene | Antibiotic Resistance |
| tnpA | IS26 | 10106-10810 | + | Transposase | |
| tniQ | In32 | 11316-12621 | - | Accessory Gene | Target Site Selection |
| tniB | In32 | 12618-13439 | - | Accessory Gene | |
| tniA | In32 | 13442-15157 | - | Transposase | None |
| RHH_6 | Tn1412 | 15370-15609 | + | Passenger Gene | Antitoxin |
| PIN_3 | Tn1412 | 15609-16040 | + | Passenger Gene | Toxin |
| tnpA | Tn1412 | 16022-19015 | + | Transposase | |
| merP | Tn1412 | 19248-19523 | - | Passenger Gene | Heavy Metal Resistance |
| merT | Tn1412 | 19536-19892 | - | Passenger Gene | Heavy Metal Resistance |
| merR | Tn1412 | 19961-20359 | + | Passenger Gene | Heavy Metal Resistance |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | Tn1412 | 309 | 41-970 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MKIGYARVST RDQKADLQVD ALKQAGCERI YQDIASGAKS ARPELDKLLA NVRPGDAVVI WKLDRLGRSL KHLVELVGEL AERKVGLQSL NDPIDTTHAQ GRLVFNLFAS LAEFERELIR ERTQAGLSAA RARGRIGGRP KGLPAKAEAT AMAAETLYRE GRLSVSAIGE KLHISKSTLY SYLRHRGVEI GAYQKSARSR DQHASAASPA EPPAAERVAT VTLRLAVVNN SKFVRGRKRA TENIERYCLE PYGMKRLDAG HYELTIPYRS DDELDKSVHD LLTEISQEAD MRNCFVEMGA WEEDTEKRW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI1 | IntI1 | In32 | 337 | 1272-2285 | - |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 1 | ||||||||||
| Sequence Family: | Class 1 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MKTATAPLPP LRSVKVLDQL RERIRYLHYS LRTEQAYVHW VRAFIRFHGV RHPATLGSSE VEAFLSWLAN ERKVSVSTHR QALAALLFFY GKVLCTDLPW LQEIGRPRPS RRLPVVLTPD EVVRILGFLE GEHRLFAQLL YGTGMRISEG LQLRVKDLDF DHGTIIVREG KGSKDRALML PESLAPSLRE QLSRARAWWL KDQAEGRSGV ALPDALERKY PRAGHSWPWF WVFAQHTHST DPRSGVVRRH HMYDQTFQRA FKRAVEQAGI TKPATPHTLR HSFATALLRS GYDIRTVQDL LGHSDVSTTM IYTHVLKVGG AGVRSPLDAL PPLTSER |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | ANT(2'')-Ia (ARO:3000230) | ANT(2'')-Ia | In32 | 177 | 2431-2964 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(2'')(ARO:3004276) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000230||Synonym=aadb | ||||||||||
| Protein Sequence: | MDTTQVTLIH KILAAADERN LPLWIGGGWA IDARLGRVTR KHDDIDLTFP GERRGELEAI VEMLGGRVME ELDYGFLAEI GDELLDCEPA WWADEAYEIA EAPQGSCPEA AEGVIAGRPV RCNSWEAIIW DYFYYADEVP PVDWPTKHIE SYRLACTSLG AEKVEVLRAA FRSRYAA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aadA (ARO:3002601) | AadA | In32 | 263 | 3022-3813 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'')(ARO:3004275) | ||||||||||
| Transposase Chemistry: | aminoglycoside nucleotidyltransferase | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002601||Synonyms: aadA1-pm, aadA, aadA1, aad(3'')(9) | ||||||||||
| Protein Sequence: | MREAVIAEVS TQLSEVVGVI ERHLEPTLLA VHLYGSAVDG GLKPHSDIDL LVTVTVRLDE TTRRALINDL LETSASPGES EILRAVEVTI VVHDDIIPWR YPAKRELQFG EWQRNDILAG IFEPATIDID LAILLTKARE HSVALVGPAA EELFDPVPEQ DLFEALNETL TLWNSPPDWA GDERNVVLTL SRIWYSAVTG KIAPKDVAAD WAMERLPAQY QPVILEARQA YLGQEEDRLA SRADQLEEFV HYVKGEITKV VGK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | ANT(2'')-Ia (ARO:3000230) | ANT(2'')-Ia | In32 | 177 | 3878-4411 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(2'')(ARO:3004276) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000230||Synonym=aadb | ||||||||||
| Protein Sequence: | MDTTQVTLIH KILAAADERN LPLWIGGGWA IDARLGRVTR KHDDIDLTFP GERRGELEAI VEMLGGRVME ELDYGFLAEI GDELLDCEPA WWADEAYEIA EAPQGSCPEA AEGVIAGRPV RCNSWEAIIW DYFYYADEVP PVDWPTKHIE SYRLACTSLG AEKVEVLRAA FRSRYAA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aadA (ARO:3002601) | AadA | In32 | 263 | 4469-5260 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'')(ARO:3004275) | ||||||||||
| Transposase Chemistry: | aminoglycoside nucleotidyltransferase | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002601||Synonyms: aadA1-pm, aadA, aadA1, aad(3'')(9) | ||||||||||
| Protein Sequence: | MREAVIAEVS TQLSEVVGVI ERHLEPTLLA VHLYGSAVDG GLKPHSDIDL LVTVTVRLDE TTRRALINDL LETSASPGES EILRAVEVTI VVHDDIIPWR YPAKRELQFG EWQRNDILAG IFEPATIDID LAILLTKARE HSVALVGPAA EELFDPVPEQ DLFEALNETL TLWNSPPDWA GDERNVVLTL SRIWYSAVTG KIAPKDVAAD WAMERLPAQY QPVILEARQA YLGQEEDRLA SRADQLEEFV HYVKGEITKV VGK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacEdelta1 (ARO:3005010) | QacEdelta1 | In32 | 115 | 5424-5771 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | subunit of the qac multidrug efflux pump||perfect match to reference sequence for ARO:3005010 (bitscore:219) | ||||||||||
| Protein Sequence: | MKGWLFLVIA IVGEVIATSA LKSSEGFTKL APSAVVIIGY GIAFYFLSLV LKSIPVGVAY AVWSGLGVVI ITAIAWLLHG QKLDAWGFVG MGLIIAAFLL ARSPSWKSLR RPTPW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | sul1 (ARO:3000410) | Sul1 | In32 | 279 | 5765-6604 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | sulfonamide resistant sul (ARO:3004238) | ||||||||||
| Transposase Chemistry: | dihydropteroate synthase | ||||||||||
| Target: | sulfonamide antibiotic (ARO:3000282)||sulfone antibiotic (ARO:3003401) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000410 | ||||||||||
| Protein Sequence: | MVTVFGILNL TEDSFFDESR RLDPAGAVTA AIEMLRVGSD VVDVGPAASH PDARPVSPAD EIRRIAPLLD ALSDQMHRVS IDSFQPETQR YALKRGVGYL NDIQGFPDPA LYPDIAEADC RLVVMHSAQR DGIATRTGHL RPEDALDEIV RFFEARVSAL RRSGVAADRL ILDPGMGFFL SPAPETSLHV LSNLQKLKSA LGLPLLVSVS RKSFLGATVG LPVKDLGPAS LAAELHAIGN GADYVRTHAP GDLRSAITFS ETLAKFRSRD ARDRGLDHA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | bla LCR-1 (ARO:3002997) | Bla LCR-1 | In32 | 260 | 7158-7940 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | LCR beta-lactamase (ARO:3002996) | ||||||||||
| Target: | penam (ARO:3000008)||cephalosporin (ARO:0000032) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002996 | ||||||||||
| Protein Sequence: | MLKSTLLAFG LFIALSARAE NQAIAKLFLR AGVDGTIVIE SLTTGQRLVH NDPRAQQRYP AASTFKVLNT LIALEEGAIS GENQIFHWNG TQYSIANWNQ DQTLDSAFKV SCVWCYQQIA LRVGALKYPA YIQQTNYGHL LEPFNGTEFW LDGSLTISAE EQVAFLRQVV ERKLPFKASS YDSLKKVMFA DENAQYRLYA KTGWATRMTP SVGWYVGYVE AKDDVWLFAL NLATRDANDL PLRTQIAKDA LKAIGAFPTK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 8246-8950 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | APH(3')-Ia (ARO:3002641) | APH(3')-Ia | Tn4352 | 271 | 9140-9955 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | APH(3') (ARO:3000126) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002641 (bitscore: 560)||Synonyms: aphA-1,apha1-1AB, APH(3')-Ic, apha7 | ||||||||||
| Protein Sequence: | MSHIQRETSC SRPRLNSNMD ADLYGYKWAR DNVGQSGATI YRLYGKPDAP ELFLKHGKGS VANDVTDEMV RLNWLTEFMP LPTIKHFIRT PDDAWLLTTA IPGKTAFQVL EEYPDSGENI VDALAVFLRR LHSIPVCNCP FNSDRVFRLA QAQSRMNNGL VDASDFDDER NGWPVEQVWK EMHKLLPFSP DSVVTHGDFS LDNLIFDEGK LIGCIDVGRV GIADRYQDLA ILWNCLGEFS PSLQKRLFQK YGIDNPDMNK LQFHLMLDEF F |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 10106-10810 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniQ | TniQ | In32 | 435 | 11316-12621 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Target Site Selection | ||||||||||
| Protein Sequence: | VRKRSTTARS AWPITPVPAS GVGSSSGN*C EASSTLAAAS GSQGRRSLIL VAQSCRRLLP DGYA*LVGTR SWA*QGR*PG HRTAAVAADG TLPAKRHRAG PTALHEFCRL GAMAAG*PR* PGSVRPGNLC VPVLGNAAKT QPQDEIGHEL ARVAAQSANP SGLPTLPE*F SEPGGTARVE ATPDAELSPA WMLAGIVLGP ARACAWLGEG RRCPAHGQ*R DCDDGPAHLA STDSRIRRVA APARPCRFVV PAAAYTAG*A EYPALALRDL RWQYPLCLGA LRPSAARRAK SVASLRDSGL IGAVADIGGS GNRHRPDRIE GVESTRRTGN AVPVRATN*V HQRPASG*AE EGTRQLLATS PQSNQRCHYR SAARLRDSPL VIRDGILWTA *P*IPGAFAR HLYQGGHSP* ISVTL*ARRP LYMP*TK*RV K*QIL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB | TniB | In32 | 273 | 12618-13439 | - |
|---|
| Class: | Accessory Gene | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Comment: | homologous to TnsC protein of Tn7 putative ATP-binding protein | ||||||||||
| Protein Sequence: | VDEYPIIDLS HLLPTAQGLA RLPADERIQR LRADRWIGYP RAVEALNRLE TLYAWPNKQR MPNLLLVGPT NNGKSMIVEK FRRTHPASAD ADQEHIPVLV VQMPSEPSVI RFYVALLAAM GAPLRPRPRL PEMEQLALAL LRKVGVRMLV IDELHNVLAG NSVNRREFLN LLRFLGNELR IPLVGVGTRD AYLAIRSDDQ LENRFEPMML PVWEANDDCC SLLASFAASL PLRRPSSIAT LDMARYLLTR SEGTIGELAH LLMAAAIAAC GER |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA | TniA | In32 | 571 | 13442-15157 | - |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MLNTRVHQKR GMGMASDARI TEHGVATLSD EAWEHARRRA EIIAPLAMLE VVGHQAADAA AQALGLSRRQ VYVLIRRARQ GSGLVTDLAP GQSGGGKGRG RLPESVERII RELLQKRFLT KQKRSLAAFH REVALVCKTH KLRVPARNTV ALRIAALDPL KTARRREGQD ASRDLQGVGG VPPAVTLPLE QIQIDHTVID LIVVDERDRQ PIGRPYLTVA IDVFTRCVLG MVVTLEAPSA VSVGLCLVHV ACDKRPWLEG LNVEMDWPMS GKPKLLYLDN AAEFKSEALR RGCEQHGIQL DYRPPGQPHY GGIVERIIGT AMQMIHDELP GTTFSNPDQR GDYDSENKAA LTLHELERWL TLAVGTYHGS VHNGLLQPPV ARWSEAVTRV GVPAVVTRAT AFLVDFLPII RRTLTRTGFV IDHIHYYADT LKPWIARRDR WPAFLIRRDP RDISRIWVLE PEGQHYLEIP YRTLSHPAVT LWEQRQALTK LRQQGREQVD ESALFRMIGQ MREIVTTAQK ATRKARRDAD RRQHLKASVP PDKPTPPETD VADPQADNLP PAKPFDQIEE W |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | RHH_6 | RHH_6 | Tn1412 | 79 | 15370-15609 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antitoxin | ||||||||||
| Function: | antitoxin||binds to DNA(IPR031914) | ||||||||||
| Sequence Family: | RHH_6 (Pfam:PF16762) | ||||||||||
| Comment: | Pfam PF16762.4 | ||||||||||
| Protein Sequence: | MNTIRWNVAV SADTDQSLRM FLASQGGGRK GDLSRFIEEA VRAHILELSA EQAKAANAHL SEAELTNAVD EALDWARKR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | PIN_3 | PIN_3 | Tn1412 | 143 | 15609-16040 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Toxin | ||||||||||
| Function: | toxin||ribonuclease activity(IPR002716) | ||||||||||
| Sequence Family: | PIN_3 (Pfam:PF13470) | ||||||||||
| Target: | single stranded RNA | ||||||||||
| Protein Sequence: | MRVVLDTNIL FSALISPHGA PDAIYRAWRA ARFEVVTSRM QLDEIRRASR YPKLQAILQP AKVGAMINNL QRAVVLERLT IEVEADDPDD SFLLAMALAG DADYLITGDR RAGLLQRGHI ERRGSSRPPC SASRCCDQCR SVF |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | Tn1412 | 997 | 16022-19015 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MPVGFLTQEQ RDGFGRYVDS PSREELERYF HLSDEDREAI QVLRGNHNRL GYAVLLTTVR FVGVLPDKPA AVPVEVLQVL CRQLAIPDPD CLQRYSDHRR WIHATDIQNR FGYRHFTDPG IGFRLSRWLY ALCWTGTDRP GVLFERATSW LFTQKVLLPG VSQLERFIAQ LRSRVEERLW FTLGRSVTEE QRLQLQDLLT VAEGNRSSRL DQLRSGPVMV SGPALIRALR RLDDVRGIGI TLPAAAHIPP SRIAALARFA NTAKVTAINR LPASRRMATL VAFALCLEAT AHDDALEVLE ALLRDLFSNA EKADKKARMR SLKDLDRSAA TLAAACKVVL DSSISDDNVR ARLFNDLPRT TLEKALEEVN ALIRPVDDVY FLALEARYRS VRRFLPDLLK HIRFGFSPAG KGVAASLEWL QLNLPRRKPE DDAPQEIVAK AWQKHITRED GSLDMGAYVF CTLDALRTAL RRRDVFVSPS WRYADPRLGL LDGAEWLAAR PIICRSLGLT IDAKTTLDAL SVELDATWLA VAARLPDNPA IQLSENTEGK TELSLGALDK LDEPCSLLQL RAAVSDLMPR VDLPEILLEI AARTGFSEAF THVSERNARA DNLVTSLCAV LLGGACNTGL EPLIRTDNPA LRRDRLSWVS QNYIRDDTLS AANAILVGAQ SQLELAQVWG GGEVASADGM RFVVPVRTVH AGPNPKYFGT GRGVTWYNLI SDQFSGLNAI TVPGTLRDSL VLLAVVLEQQ TELQPTQIMT DTGAYSDVVF GLFRLLGYHF SPRLADVGGT RFWRTRPDAD YGKLNGLARQ SVKLDLIAEH WDDLLRLAGS LKLGRVPATG IMRTLQTGDR PTRLAQALAE FGRIEKTLHT LTYIDDESKR RATLTQLNRG EGRHSLARAV FHGKRGELRQ RYREGQEDQL GALGLVVNII VLWNTLYMTA AVERLKQHGY PVLEEDLARL SPLIYEHINM LGRYSFAVPE EVARGELRPL RNPDDDL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merP | MerP | Tn1412 | 91 | 19248-19523 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Comment: | periplasmic binding protein | ||||||||||
| Protein Sequence: | MRKLLIAVLF ALPFVALAAP PKTVTLDVQN MTCGLCPITV KKSLEKVSGV SDVQVNFDQK TATVTYDPDK AQPEALTEAT ANAGYPSTVQ K |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merT | MerT | Tn1412 | 118 | 19536-19892 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Comment: | transport protein | ||||||||||
| Protein Sequence: | MGMGMQLTGK GSLVASSLTA IGASVCCVGP LVLLALGVGG TWVGALTMME PLRPLFIGLT LLFLGLAFRK LYLVPQVCTP GTPCADPRTL VRQRLVFWIV SVLLLGLLAV PWLAPLFY |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | merR | MerR | Tn1412 | 132 | 19961-20359 | + |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Mercury | ||||||||||
| Comment: | regulatory protein | ||||||||||
| Protein Sequence: | MATELTTGKL ADAAGVNVET IRYYQRRGLL DEPAKPLGGH RRYPVDMVKR LRFIKRAQAL GFTLSEVGGL LTLDESCACA ETRARAARKL ALIEQKMADL VVMQQLLGEL VQQCDAGDGG TICPIIEALI RE |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| In32-L36547 | In32 | integron | 1070-15262 | + | 14193 |
| Tn4352-M20306 | Tn4352 | transposon | 8183-10862 | + | 2680 |
| IS26-MH257753 | IS26 | insertion sequence | 8183-9002 | + | 820 |
| IS26-MH257753 | IS26 | insertion sequence | 10043-10862 | + | 820 |
| Name | Associated TE | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|---|
| IRi | In32 | 1070-1102 | + | 33 | TGTCGTTTTC AGAAGACGGC TGCACTGAAC GTC |
| repeat i1 | In32 | 1078-1096 | + | 19 | TCAGAAGACG GCTGCACTG |
| repeat i2 | In32 | 1101-1119 | + | 19 | TCAGAAGCCG ACTGCACTA |
| repeat i3 | In32 | 1143-1161 | + | 19 | TCAGGCAACG ACGGGCTGC |
| repeat i4 | In32 | 1171-1189 | + | 19 | TCAGCGGACG CAGGGAGGA |
| IRi | In32 | 6981-7013 | + | 33 | TGTCGTTTTC AGAAGACGGC TGCACTGAAC GTC |
| repeat i1 | In32 | 6989-7007 | + | 19 | TCAGAAGACG GCTGCACTG |
| repeat i2 | In32 | 7012-7030 | + | 19 | TCAGAAGCCG ACTGCACTA |
| repeat i3 | In32 | 7054-7072 | + | 19 | TCAGGCAACG ACGGGCTGC |
| IR | IS26 | 8183-8196 | + | 14 | GGCACTGTTG CAAA |
| IR | IS26 | 8989-9002 | - | 14 | GGCACTGTTG CAAA |
| IR | IS26 | 10043-10056 | + | 14 | GGCACTGTTG CAAA |
| IR | IS26 | 10849-10862 | - | 14 | GGCACTGTTG CAAA |
| repeat t4 | In32 | 15135-15153 | - | 19 | TCAATACTCG TGTGCACCA |
| repeat t1 | In32 | 15236-15254 | - | 19 | TCAGAAGACG ACTGCACCA |
| IRt | In32 | 15238-15262 | - | 25 | TGTCGTTTTC AGAAGACGAC TGCAC |
1. Partridge SR, Brown HJ, Stokes HW, Hall RM.
Transposons Tn1696 and Tn21 and their integrons In4 and In2 have independent origins.. Antimicrob Agents Chemother. 2001 Apr;45(4):1263-70. doi: 10.1128/AAC.45.4.1263-1270.2001. PubMed ID:
11257044
.
2. Partridge SR, Recchia GD, Stokes HW, Hall RM. Family of class 1 integrons related to In4 from Tn1696.. Antimicrob Agents Chemother. 2001 Nov;45(11):3014-20. doi: 10.1128/AAC.45.11.3014-3020.2001. PubMed ID: 11600350 .
3. Szuplewska M, Ludwiczak M, Lyzwa K, Czarnecki J, Bartosik D. Mobility and generation of mosaic non-autonomous transposons by Tn3-derived inverted-repeat miniature elements (TIMEs).. PLoS One. 2014 Aug 14;9(8):e105010. doi: 10.1371/journal.pone.0105010. eCollection 2014. PubMed ID: 25121765 .
2. Partridge SR, Recchia GD, Stokes HW, Hall RM. Family of class 1 integrons related to In4 from Tn1696.. Antimicrob Agents Chemother. 2001 Nov;45(11):3014-20. doi: 10.1128/AAC.45.11.3014-3020.2001. PubMed ID: 11600350 .
3. Szuplewska M, Ludwiczak M, Lyzwa K, Czarnecki J, Bartosik D. Mobility and generation of mosaic non-autonomous transposons by Tn3-derived inverted-repeat miniature elements (TIMEs).. PLoS One. 2014 Aug 14;9(8):e105010. doi: 10.1371/journal.pone.0105010. eCollection 2014. PubMed ID: 25121765 .