Mobile element type:
Integron
Name:
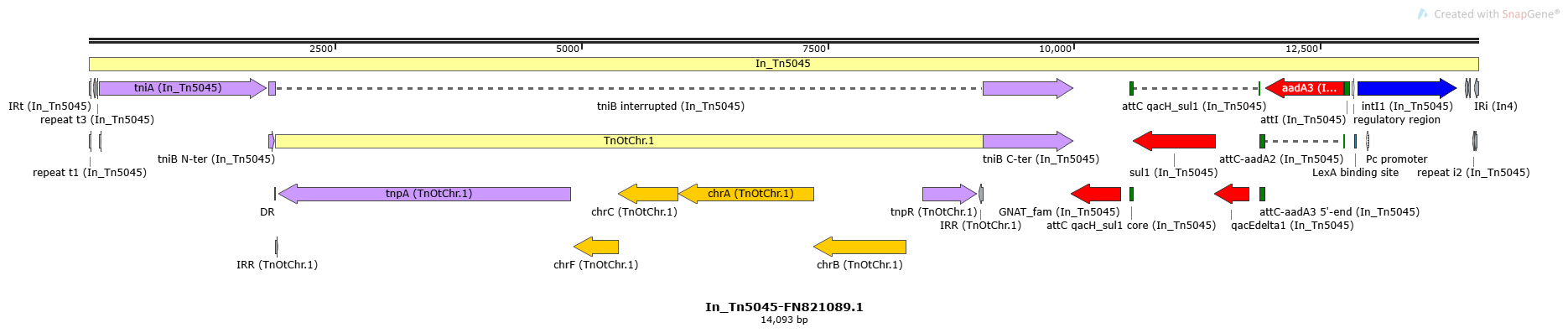
In_Tn5045
Synonyms:
Accession:
In_Tn5045-FN821089.1
Family:
Tn402
Group:
Class 1
First isolate:
Yes
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Pseudomonas sp. Tik3
Date of Isolation:
2011
Country:
Russia
Molecular Source:
plasmid
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRt | 1-33 | + | 33 | TGTCATTTTC AGAAGACGAC TGCACCAGTT GAT |
| repeat t1 | 9-27 | + | 19 | TCAGAAGACG ACTGCACCA |
| repeat t2 | 49-67 | - | 19 | TCAGGAGCTG GCTGCACAA |
| repeat t3 | 78-97 | + | 20 | TCAGAAGTGA TCTGCACCAA |
| repeat t4 | 110-128 | + | 19 | TCAATACTCG TGTGCACCA |
| repeat i4 | 13974-13992 | - | 19 | TCAGCGGACG CAGGGAGGA |
| repeat i3 | 14002-14020 | - | 19 | TCAGGCAACG ACGGGCTGC |
| repeat i2 | 14044-14062 | - | 19 | TCAGAAGCCG ACTGCACTA |
| repeat i1 | 14067-14085 | - | 19 | TCAGAAGACG GCTGCACTG |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| attC qacH_sul1 | In_Tn5045 | 10569-10602 , 11880-11886 | + | 41 | CCGCTAGCGG GCGGCCGGAA GGTGAATGCT AGGCATCTAA C |
| attC qacH_sul1 core | In_Tn5045 | 10569-10602 | + | 34 | CCGCTAGCGG GCGGCCGGAA GGTGAATGCT AGGC |
| attC-aadA2 | In_Tn5045 | 11886-11939 , 12736-12741 | + | 60 | CGCCGGAGTT AAGCCGCCGC GCGTAGCGCG GTCGGCTTGA ACGAATTGTT AGACGTCTAA |
| attC-aadA3 5'-end | In_Tn5045 | 11886-11939 | + | 54 | CGCCGGAGTT AAGCCGCCGC GCGTAGCGCG GTCGGCTTGA ACGAATTGTT AGAC |
| attI | In_Tn5045 | 12742-12797 | + | 56 | CTTTGTTTTA GGGCGACTGC CCTGCTGCGT AACATCGTTG CTGCTCCATA ACATCA |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tniA | In_Tn5045 | 106-1821 | + | Transposase | |
| tniB N-ter | In_Tn5045 | 1824-1892 | + | Transposase | |
| tnpA | TnOtChr.1 | 1926-4892 | - | Transposase | |
| chrF | TnOtChr.1 | 4923-5375 | - | Passenger Gene | Heavy Metal Resistance |
| chrC | TnOtChr.1 | 5372-5980 | - | Passenger Gene | Heavy Metal Resistance |
| chrA | TnOtChr.1 | 5990-7357 | - | Passenger Gene | Heavy Metal Resistance |
| chrB | TnOtChr.1 | 7354-8292 | - | Passenger Gene | Heavy Metal Resistance |
| tnpR | TnOtChr.1 | 8463-9029 | + | Accessory Gene | Resolvase |
| tniB C-ter | In_Tn5045 | 9073-10000 | + | Transposase | |
| GNAT_fam | In_Tn5045 | 9969-10469 | - | Passenger Gene | Antibiotic Resistance |
| sul1 (ARO:3000410) | In_Tn5045 | 10597-11436 | - | Passenger Gene | Antibiotic Resistance |
| qacEdelta1 (ARO:3005010) | In_Tn5045 | 11430-11777 | - | Passenger Gene | Antibiotic Resistance |
| aadA3 (ARO:3002603) | In_Tn5045 | 11941-12732 | - | Passenger Gene | Antibiotic Resistance |
| intI1 | In_Tn5045 | 12878-13891 | + | Integron Integrase | Class 1 |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA | TniA | In_Tn5045 | 571 | 106-1821 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | identical to tniA (Tn1721) | ||||||||||
| Protein Sequence: | MLNTRVHQSE VSMATDTPRI PEQGVATLPD EAWERARRRA EIISPLAQSE TVGHEAADMA AQALGLSRRQ VYVLIRRARQ GSGLVTDLVP GQSGGGKGKG RLPEPVERVI HELLQKRFLT KQKRSLAAFH REVTQVCKAQ KLRVPARNTV ALRIASLDPR KVIRRREGQD AARDLQGVGG EPPAVTAPLE QVQIDHTVID LIVVDDRDRQ PIGRPYLTLA IDVFTRCVLG MVVTLEAPSA VSVGLCLVHV ACDKRPWLEG LNVEMDWQMS GKPLLLYLDN AAEFKSEALR RGCEQHGIRL DYRPLGQPHY GGIVERIIGT AMQMIHDELP GTTFSNPDQR GDYDSENKAA LTLRELERWL TLAVGTYHGS VHNGLLQPPA ARWAEAVARV GVPAVVTRAT SFLVDFLPIL RRTLTRTGFV IDHIHYYADA LKPWIARRER WPSFLIRRDP RDISRIWVLE PEGQHYLEIP YRTLSHPAVT LWEQRQALAK LRQQGREQVD ESALFRMIGQ MREIVTSAQK ATRKARRDAD RRQHLKTSAR PDKPVPPDTD IADPQADNLP PAKPFDQIEE W |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB N-ter | TniB N-ter | In_Tn5045 | 23 | 1824-1892 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | disrupted by TnOtChr.1 | ||||||||||
| Protein Sequence: | VDEYPIIDLS HLLPAAQGLA RLP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | TnOtChr.1 | 988 | 1926-4892 | - |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MPRRSILSAA ERESLLALPD TMEDLIRYYT FSESDLSIIR QRRGPANLLG FAVQLCYMRY PGMMLAVDTE PFPPLLRLVA TQLKVPPEAW ADYGQRAETR REHLLELQSI FGFQAFATRH YRPSVHSLDE LAWQTDKGIV LATELIEGLR QKSVLLPSPG VIERICAEAI TRANRRIYEA LSEPLTDPHR HRLDDLLKRR EFGKTTWLAW LRQSPVKPNS RHMLEHIERL KAWQALDLPT GIERLLHQNR LLKIAREGGQ MTPADLAKFE PQRRYATLVA LAIEGMATVI DEIIDLHDRI LGKLFNAAKH KHQQQFQASG KAINAKVRLY GRIGQALIDA KQSGGNPFAA IEAVMSWDAF AQSVTEAQKL AQPDDFDFLH RIGESYATLR RYAPEFLAVL KLRAAPAAKG VLDAVDVLRG MNTDNVRKVP TDAPTDFIKP RWQKLVMTDA GIDRRYYELC ALSELKNSLR SGDIWVQGSR QFKDFEDYLV PHEKFASLKQ GSELPLDVAT DCDQYLYERL TLLEAQLATV NRMAAANDLP DAIITESGLK ITPLDAAVPD TAQALIDQTA MILPHVKITE LLLEVDEWTG FTRYFTHLKS GDLAKDKNLL LTTILADAIN LGLSKMAESC PGTTYAKLAW LQAWHIRDET YSTALAELVN TQFRHPFACH WGDGTTSSSD GQNFRTGSKA KSTGHINPKY GSSPGRTFYT HISDQYAPFH AKVVNVGVRD STYVLDGLLY HESDLRIEEH YTDTAGFTDH VFALMHLLGF RFAPRIRDLG DTKLYLPKGD TAYEALKPMV GGTLNIKHVR AHWDEILRLA TSIKHGTVTA SLMLRKLGSY PRQNGLAVAL RELGRIERTL FILDWLQSVE LRRRVHAGLN KGEARNALAR AVFFNRLGEI RDRSFEQQRY RASGLNLVTA AVVLWNTVYL ERAAHALRGN GLGIDDALLQ YLSPLGWEHI NLTGDYLWRS SAKIGAGKFR PLRPLQPP |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | chrF | ChrF | TnOtChr.1 | 150 | 4923-5375 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Chromate | ||||||||||
| Protein Sequence: | MKWITRERPK IDRIACPWMV TRFIDTHAEF LYVPAGDVLR IAQETGAVPY DIPGVELSHD GELCSFDAFL AKYAMDDPAL KQLAVIVRGA DTSRLDLTPQ STGLYALSLG LSQNFSDDHE MLKHGMVMYD AFYAWCKHCQ SETHNWPPKM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | chrC | ChrC | TnOtChr.1 | 202 | 5372-5980 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Chromate | ||||||||||
| Comment: | superoxide dismutase unkown function | ||||||||||
| Protein Sequence: | MSFDIKPLPF PASTLNGLSE KLIASHHENN YSGAVKRLNA IHTKLGTLDF TQEAGFLLNG LKREELIAYN SMLLHEIYFD SLGGNGVLPV GPLCEAIERD FGSVERWQAE FSAMGKALGG GSGWVLLVQS ARDGKLSNQW AADHCHTLAG ALPILALDMY EHAYHIDYGA KAGAYVDAFM ANIDWQRAAI RWASHQPSGG LS |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | chrA | ChrA | TnOtChr.1 | 455 | 5990-7357 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Chromate | ||||||||||
| Comment: | chromate efflux pump | ||||||||||
| Protein Sequence: | MTQTVVTKEA VAVPPSPQVV SFWQAFRLWL KIGFIGFGGP AGQIAIMHRE LVEQRRWISE RRFLHALNFC MLLPGPEGQQ LATYMGWLMH RTWGGIVAGV LFVLPSLFFL IALSWIYIAF GDTPLVSGLF YGIKPAITAV VVQAVHRIGS RALKNGLMWA IAAGAFVAIF AMNVPFPAIV AGAALIGYIG GRVTPDKFRA GGAHRAADKS YGPALIDDNT PIPSHALFRW SGSLRVAAMG CLLWLIPMAF LLGTYGWDHT LTQMGWFFTK AALLTFGGAY AVLPYIYQGA VGHYSWLTPT QMVDGLALGE ANPGPLIMVV AFVGFVGAYV QALFGPDMLF VAGAVAAALV TWFTFLPSFI FILAGGPFVE STRGDLRFTA PLTAITAAVV GVILNLAVFF AYHVLWPKGL AGLFDWVSAL ITIGAAVALF RFKRNVIHVI AACAVLGLIL KTLIL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | chrB | ChrB | TnOtChr.1 | 312 | 7354-8292 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Heavy Metal Resistance | ||||||||||
| Target: | Chromate | ||||||||||
| Comment: | chromate-sensitive regulator of chrBACF operon exclusively activated by Cr(VI) | ||||||||||
| Protein Sequence: | MNLLSLILSL PTENATVRQR TWRALKASGA AVLRDGVYLM PDRDECRAVL DNLASDVREG GGVAHVLRME DPEGVNFVAL FDRSNDFAAL LVDVHHLRQT LTLDTVQDVL RQVRKLRKSF TTLVEIDFYP GEAQRQADSA LCELEQACAR TLSPDEPHAV EGTITRLDRL DYQARTWATR ARPWVDRLAS AWLIRRFIDP QARILWLATP ADCPPDALGF DFDGATFSHV GSRVTFEVLA ASFGLEQPAI TRIGLVVHYL DVGGIQPPEA TGIESVLAGL RETVDHDDQL LAIASTVFDG LLASFEKGTL TV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpR | TnpR | TnOtChr.1 | 188 | 8463-9029 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Subclass: | Resolvase | ||||||||||
| Sequence Family: | Serine Site-Specific Recombinase | ||||||||||
| Transposase Chemistry: | Serine | ||||||||||
| Protein Sequence: | MRIGYARVST QEQDNQAQIS ALQSAGCELI FQEKASGGRW DRPELHRLLG HLRKADVVVV WKLDRLSRSL KDLLLTLEKI EEAGAGFQSL TESIDTTTPA GRMMMQIVGS FAEFERAMLR ERTRHGLEAA RKDGRVGGRR PKLTQQQQKE IVALITSGQK TGADAARLFR VHPSTVVRLL AKHRQGPG |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB C-ter | TniB C-ter | In_Tn5045 | 308 | 9073-10000 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | disrupted by TnOtChr.1 | ||||||||||
| Protein Sequence: | PADERIQRLR ADRWIGYPRA VEALNRLEAL YAWPNKQRMP NLLLVGPTNN GKSMIVEKFR RTHPASSDAD QEHIPVLVVQ MPSEPSVIRF YVALLAAMGA PLRPRPRLPE MEQLALALLR KVGVRMLVID ELHNVLAGNS VNRREFLNLL RFLGNELRIP LVGVGTRDAY LAIRSDDQLE NRFEPMMLPV WEANDDCCSL LASFAASLPL RRPSPIATLD MARYLLTRSE GTIGELAHLL MAAAIVAVES GEEAINHRTL SMACRQPLAQ PRHRGRTASD LEATGPSHPD MDLDARTDVR FRVLGVLR |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | GNAT_fam | GNAT_fam | In_Tn5045 | 166 | 9969-10469 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Sequence Family: | Acetyltransf_1 (Pfam:PF00583) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | putative acetyltransferase ADU64769.1 | ||||||||||
| Protein Sequence: | MDSEEPPNVR VACSGDIDEV VRLMHDAAAW MSAKGTPAWD VARIDRTFAE TFVLRSELLV ASCSDGIVGC CTLSAEDPEF WPDALKGEAA YLHKLAVRRT HAGRGVSSAL IEACRHAART QGCAKLRLDC HPNLRGLYER LGFTHVDTFN PGWDPTFIAE RLELEI |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | sul1 (ARO:3000410) | Sul1 | In_Tn5045 | 279 | 10597-11436 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | sulfonamide resistant sul (ARO:3004238) | ||||||||||
| Transposase Chemistry: | dihydropteroate synthase | ||||||||||
| Target: | sulfonamide antibiotic (ARO:3000282)||sulfone antibiotic (ARO:3003401) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000410 | ||||||||||
| Protein Sequence: | MVTVFGILNL TEDSFFDESR RLDPAGAVTA AIEMLRVGSD VVDVGPAASH PDARPVSPAD EIRRIAPLLD ALSDQMHRVS IDSFQPETQR YALKRGVGYL NDIQGFPDPA LYPDIAEADC RLVVMHSAQR DGIATRTGHL RPEDALDEIV RFFEARVSAL RRSGVAADRL ILDPGMGFFL SPAPETSLHV LSNLQKLKSA LGLPLLVSVS RKSFLGATVG LPVKDLGPAS LAAELHAIGN GADYVRTHAP GDLRSAITFS ETLAKFRSRD ARDRGLDHA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacEdelta1 (ARO:3005010) | QacEdelta1 | In_Tn5045 | 115 | 11430-11777 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | subunit of the qac multidrug efflux pump||perfect match to reference sequence for ARO:3005010 (bitscore:219) | ||||||||||
| Protein Sequence: | MKGWLFLVIA IVGEVIATSA LKSSEGFTKL APSAVVIIGY GIAFYFLSLV LKSIPVGVAY AVWSGLGVVI ITAIAWLLHG QKLDAWGFVG MGLIIAAFLL ARSPSWKSLR RPTPW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aadA3 (ARO:3002603) | AadA3 | In_Tn5045 | 263 | 11941-12732 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'') (ARO:3004275) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002603 (bitscore: 522) | ||||||||||
| Protein Sequence: | MREAVTIEIS NQLSEVLSVI ERHLESTLLA VHLYGSAVDG GLKPYSDIDL LVTVAVKLDE TTRRALLNDL MEASAFPGES ETLRAIEVTL VVHDDIIPWR YPAKRELQFG EWQRNDILAG IFEPAMIDID LAILLTKARE HSVALVGPAA EEFFDPVPEQ DLFEALRETL KLWNSQPDWA GDERNVVLTL SRIWYSAITG KIAPKDVAAD WAIKRLPAQY QPVLLEAKQA YLGQKEDHLA SRADHLEEFI RFVKGEIIKS VGK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI1 | IntI1 | In_Tn5045 | 337 | 12878-13891 | + |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 1 | ||||||||||
| Sequence Family: | Class 1 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MKTATAPLPP LRSVKVLDQL RERIRYLHYS LRTEQAYVHW VRAFIRFHGV RHPATLGSSE VEAFLSWLAN ERKVSVSTHR QALAALLFFY GKVLCTDLPW LQEIGRPRPS RRLPVVLTPD EVVRILGFLE GEHRLFAQLL YGTGMRISEG LQLRVKDLDF DHGTIIVREG KGSKDRALML PESLAPSLRE QLSRARAWWL KDQAEGRSGV ALPDALERKY PRAGHSWPWF WVFAQHTHST DPRSGVVRRH HMYDQTFQRA FKRAVEQAGI TKPATPHTLR HSFATALLRS GYDIRTVQDL LGHSDVSTTM IYTHVLKVGG AGVRSPLDAL PPLTSER |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| TnOtChr.1-FN821089.1 | TnOtChr.1 | transposon | 1893-9072 | + | 7180 |
| Name | Associated TE | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|---|
| IRR | TnOtChr.1 | 1893-1930 | + | 38 | GGGGTCGTCT CAGAAAACGG ACCGCAAAGT ACGCTAAG |
| IRR | TnOtChr.1 | 9035-9072 | - | 38 | GGGGTCGTCT CAGAAAACGG ACCGCAAAGT ACGCTAAG |
1. Petrova M, Gorlenko Z, Mindlin S.
Tn5045, a novel integron-containing antibiotic and chromate resistance transposon isolated from a permafrost bacterium.. Res Microbiol. 2011 Apr;162(3):337-45. doi: 10.1016/j.resmic.2011.01.003. Epub 2011 Jan 22. PubMed ID:
21262357
.