Mobile element type:
Integron
Name:
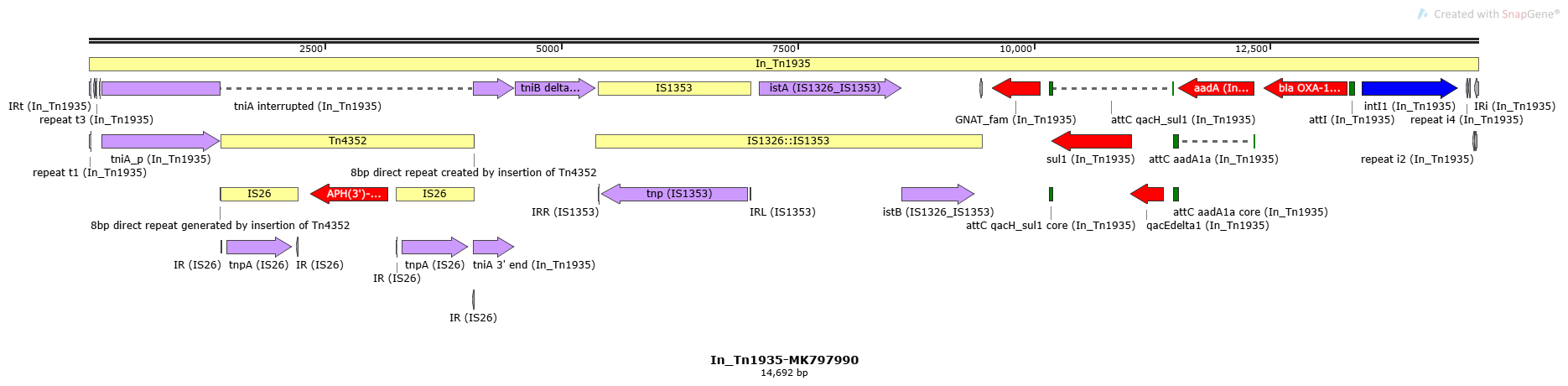
In_Tn1935
Synonyms:
Accession:
In_Tn1935-MK797990
Family:
Tn402
Group:
Class 1
First isolate:
Yes
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Salmonella enterica subsp. enterica serovar Wien ZM3
Date of Isolation:
2019
Country:
Molecular Source:
Tn1935 in plasmid pZM3
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| IRt | 1-33 | + | 33 | TGTCATTTTC AGAAGACGAC TGCACCAGTT GAT |
| repeat t1 | 9-27 | + | 19 | TCAGAAGACG ACTGCACCA |
| repeat t2 | 49-67 | - | 19 | TCAGGAGCTG GCTGCACAA |
| repeat t3 | 78-97 | + | 20 | TCAGAAGTGA TCTGCACCAA |
| repeat t4 | 110-128 | + | 19 | TCAATACTCG TGTGCACCA |
| repeat i4 | 14573-14591 | - | 19 | TCAGCGGACG CAGGGAGGA |
| repeat i3 | 14601-14619 | - | 19 | TCAGGCAACG ACGGGCTGC |
| repeat i2 | 14643-14661 | - | 19 | TCAGAAGCCG ACTGCACTA |
| IRi | 14660-14692 | - | 33 | TGTCGTTTTC AGAAGACGGC TGCACTGAAC GTC |
| repeat i1 | 14666-14684 | - | 19 | TCAGAAGACG GCTGCACTG |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| attC qacH_sul1 | In_Tn1935 | 10161-10194 , 11472-11478 | + | 41 | CCGCTAGCGG GCGGCCGGAA GGTGAATGCT AGGCATCTAA C |
| attC qacH_sul1 core | In_Tn1935 | 10161-10194 | + | 34 | CCGCTAGCGG GCGGCCGGAA GGTGAATGCT AGGC |
| attC aadA1a core | In_Tn1935 | 11478-11531 | + | 54 | CGCTTGAGTT AAGCCGCGCC GCGAAGCGGC GTCGGCTTGA ACGAATTGTT AGAC |
| attI | In_Tn1935 | 13338-13393 | + | 56 | CTTTGTTTTA GGGCGACTGC CCTGCTGCGT AACATCGTTG CTGCTCCATA ACATCA |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tniA interrupted | In_Tn1935 | 142-1394 , 4073-4509 | + | Transposase | |
| tniA_p | In_Tn1935 | 142-1393 | + | Transposase | |
| tnpA | IS26 | 1457-2161 | + | Transposase | |
| APH(3')-Ia (ARO:3002641) | Tn4352 | 2351-3166 | - | Passenger Gene | Antibiotic Resistance |
| tnpA | IS26 | 3317-4021 | + | Transposase | |
| tniA 3' end | In_Tn1935 | 4069-4509 | + | Transposase | |
| tniB delta1 | In_Tn1935 | 4512-5372 | + | Accessory Gene | |
| tnp | IS1353 | 5423-6967 | - | Transposase | |
| GNAT_fam | In_Tn1935 | 9561-10061 | - | Passenger Gene | Antibiotic Resistance |
| sul1 (ARO:3000410) | In_Tn1935 | 10189-11028 | - | Passenger Gene | Antibiotic Resistance |
| qacEdelta1 (ARO:3005010) | In_Tn1935 | 11022-11369 | - | Passenger Gene | Antibiotic Resistance |
| aadA (ARO:3002601) | In_Tn1935 | 11533-12324 | - | Passenger Gene | Antibiotic Resistance |
| bla OXA-1 (ARO:3001396) | In_Tn1935 | 12437-13312 | - | Passenger Gene | Antibiotic Resistance |
| intI1 | In_Tn1935 | 13477-14490 | + | Integron Integrase | Class 1 |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA interrupted | TniA interrupted | In_Tn1935 | 563 | 142-1394 , 4073-4509 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | can be extended upstream by 12 amino acids||TnAph1 insertion | ||||||||||
| Protein Sequence: | MATDTPRIPE QGVATLPDEA WERARRRAEI ISPLAQSETV GHEAADMAAQ ALGLSRRQVY VLIRRARQGS GLVTDLVPGQ SGGGKGKGRL PEPVERVIHE LLQKRFLTKQ KRSLAAFHRE VTQVCKAQKL RVPARNTVAL RIASLDPRKV IRRREGQDAA RDLQGVGGEP PAVTAPLEQV QIDHTVIDLI VVDDRDRQPI GRPYLTLAID VFTRCVLGMV VTLEAPSAVS VGLCLVHVAC DKRPWLEGLN VEMDWQMSGK PLLLYLDNAA EFKSEALRRG CEQHGIRLDY RPLGQPHYGG IVERIIGTAM QMIHDELPGT TFSNPDQRGD YDSENKAALT LRELERWLTL AVGTYHGSVH NGLLQPPAAR WAEAVARVGV PAVVTRATSF LVDFLPILRR TLTRTGFVID HIHYYADGRR CAQAVDCAA* TLAVLSDPAR SARHQPYLGP GTGGTALPGN SLPYLVASGC HPLGTTAGAG ETAAARARTG G*VGAVPHDR PDA*DCDQRA EGHTQGAA*R GSPPAPQDIS SAGQARSAGY GYCRPAGRQL ATRQTVRPD* GVV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA_p | TniA_p | In_Tn1935 | 417 | 142-1393 | + |
|---|
| Class: | Transposase | ||||||||||
| Function: | integrase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | could be extanded 12 amino acids upstream||Contains the first 429 amino acids of tniA (In2)||probably truncated by insertion of IS26 | ||||||||||
| Protein Sequence: | MATDTPRIPE QGVATLPDEA WERARRRAEI ISPLAQSETV GHEAADMAAQ ALGLSRRQVY VLIRRARQGS GLVTDLVPGQ SGGGKGKGRL PEPVERVIHE LLQKRFLTKQ KRSLAAFHRE VTQVCKAQKL RVPARNTVAL RIASLDPRKV IRRREGQDAA RDLQGVGGEP PAVTAPLEQV QIDHTVIDLI VVDDRDRQPI GRPYLTLAID VFTRCVLGMV VTLEAPSAVS VGLCLVHVAC DKRPWLEGLN VEMDWQMSGK PLLLYLDNAA EFKSEALRRG CEQHGIRLDY RPLGQPHYGG IVERIIGTAM QMIHDELPGT TFSNPDQRGD YDSENKAALT LRELERWLTL AVGTYHGSVH NGLLQPPAAR WAEAVARVGV PAVVTRATSF LVDFLPILRR TLTRTGFVID HIHYYAD |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 1457-2161 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | APH(3')-Ia (ARO:3002641) | APH(3')-Ia | Tn4352 | 271 | 2351-3166 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | APH(3') (ARO:3000126) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002641 (bitscore: 560)||Synonyms: aphA-1,apha1-1AB, APH(3')-Ic, apha7 | ||||||||||
| Protein Sequence: | MSHIQRETSC SRPRLNSNMD ADLYGYKWAR DNVGQSGATI YRLYGKPDAP ELFLKHGKGS VANDVTDEMV RLNWLTEFMP LPTIKHFIRT PDDAWLLTTA IPGKTAFQVL EEYPDSGENI VDALAVFLRR LHSIPVCNCP FNSDRVFRLA QAQSRMNNGL VDASDFDDER NGWPVEQVWK EMHKLLPFSP DSVVTHGDFS LDNLIFDEGK LIGCIDVGRV GIADRYQDLA ILWNCLGEFS PSLQKRLFQK YGIDNPDMNK LQFHLMLDEF F |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnpA | TnpA | IS26 | 234 | 3317-4021 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MNPFKGRHFQ RDIILWAVRW YCKYGISYRE LQEMLAERGV NVDHSTIYRW VQRYAPEMEK RLRWYWRNPS DLCPWHMDET YVKVNGRWAY LYRAVDSRGR TVDFYLSSRR NSKAAYRFLG KILNNVKKWQ IPRFINTDKA PAYGRALALL KREGRCPSDV EHRQIKYRNN VIECDHGKLK RIIGATLGFK SMKTAYATIK GIEVMRALRK GQASAFYYGD PLGEMRLVSR VFEM |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA 3' end | TniA 3' end | In_Tn1935 | 146 | 4069-4509 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | Contains the last 146 amino acids of tniA (In2)||probably truncated by insertion of IS26 | ||||||||||
| Protein Sequence: | MPADALKPWI ARRERWPSFL IRRDPRDISR IWVLEPEGQH YLEIPYRTLS HPAVTLWEQR QALAKLRQQG REQVDESALF RMIGQMREIV TSAQKATRKA RRDADRRQHL KTSARPDKPV PPDTDIADPQ ADNLPPAKPF DQIEEW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB delta1 | TniB delta1 | In_Tn1935 | 286 | 4512-5372 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Function: | probable ATP-binding protein. | ||||||||||
| Comment: | probably truncated by insertion of IS1326::IS1353 | ||||||||||
| Protein Sequence: | MDEYPIIDLS HLLPAAQGLA RLPADERIQR LRADRWIGYP RAVEALNRLE ALYAWPNKQR MPNLLLVGPT NNGKSMIVEK FRRTHPASSD ADQEHIPVLV VQMPSEPSVI RFYVALLAAM GAPLRPRPRL PEMEQLALAL LRKVGVRMLV IDELHNVLAG NSVNRREFLN LLRFLGNELR IPLVGVGTRD AYLAIRSDDQ LENRFEPMML PVWEANDDCC SLLASFAASL PLRRPSPIAT LDMARYLLTR SEGTIGELAH LLMAAAIVAV ESGEEAINHR TLSMAC |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tnp | Tnp | IS1353 | 514 | 5423-6967 | - |
|---|
| Class: | Transposase | ||||||||||
| Function: | transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Protein Sequence: | MYSYEDRLRA VRLYLKLGRR MSATLRQLGY PTKNSLKAWL AEFERNQDLR RGYQRIKRQY TDEQKQRAVD HYIEQGYCLS HTIRSLGYPS REALRAWIRD LRPEFARTVV GSSAPTVARS RLEKQQAVIA LNLRVGSAKD VADTVGVSRP TLYNWQHRLL GKVPLKPMTK KKGDTSLEQR HEALLRELAE LESQNQRLRM ENAILEKASE LIKKDMGINP LELTSREKTK VVDALRVTFP LANLLCGLKL ARSTYFYQRL RQTRPDKYTQ VREVIRTIFE DNYRCYGYRR IDSALRLGGM RVSEKVVRRL MAQERLVVRT PRRRRFSAYA GDPTPAVPNL LNRDFHASAP NTKWLTDLTE IHIPAGKVYV SPIVDCFDGL VVAWNIGTSP DANLVNTMLD HAVRTLRPGE HPVIHSDRGS HYRWPAWIRR TENAQLTRSM SKKGCSPDNA ACEGFFGRLK TELIYPRNWQ HVTLKDLMTR IDAYIHWYNE RRIKVSLGGR SPIEYRHAVG LMSV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | GNAT_fam | GNAT_fam | In_Tn1935 | 166 | 9561-10061 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Sequence Family: | Acetyltransf_1 (Pfam:PF00583) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | putative acetyltransferase ADU64769.1 | ||||||||||
| Protein Sequence: | MDSEEPPNVR VACSGDIDEV VRLMHDAAAW MSAKGTPAWD VARIDRTFAE TFVLRSELLV ASCSDGIVGC CTLSAEDPEF WPDALKGEAA YLHKLAVRRT HAGRGVSSAL IEACRHAART QGCAKLRLDC HPNLRGLYER LGFTHVDTFN PGWDPTFIAE RLELEI |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | sul1 (ARO:3000410) | Sul1 | In_Tn1935 | 279 | 10189-11028 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | sulfonamide resistant sul (ARO:3004238) | ||||||||||
| Transposase Chemistry: | dihydropteroate synthase | ||||||||||
| Target: | sulfonamide antibiotic (ARO:3000282)||sulfone antibiotic (ARO:3003401) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000410 | ||||||||||
| Protein Sequence: | MVTVFGILNL TEDSFFDESR RLDPAGAVTA AIEMLRVGSD VVDVGPAASH PDARPVSPAD EIRRIAPLLD ALSDQMHRVS IDSFQPETQR YALKRGVGYL NDIQGFPDPA LYPDIAEADC RLVVMHSAQR DGIATRTGHL RPEDALDEIV RFFEARVSAL RRSGVAADRL ILDPGMGFFL SPAPETSLHV LSNLQKLKSA LGLPLLVSVS RKSFLGATVG LPVKDLGPAS LAAELHAIGN GADYVRTHAP GDLRSAITFS ETLAKFRSRD ARDRGLDHA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacEdelta1 (ARO:3005010) | QacEdelta1 | In_Tn1935 | 115 | 11022-11369 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | subunit of the qac multidrug efflux pump||perfect match to reference sequence for ARO:3005010 (bitscore:219) | ||||||||||
| Protein Sequence: | MKGWLFLVIA IVGEVIATSA LKSSEGFTKL APSAVVIIGY GIAFYFLSLV LKSIPVGVAY AVWSGLGVVI ITAIAWLLHG QKLDAWGFVG MGLIIAAFLL ARSPSWKSLR RPTPW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | aadA (ARO:3002601) | AadA | In_Tn1935 | 263 | 11533-12324 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | ANT(3'') (ARO:3004275) | ||||||||||
| Transposase Chemistry: | aminoglycoside nucleotidyltransferase | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3002601||Synonyms: aadA1-pm aadA, aadA1, aad(3'')(9) | ||||||||||
| Protein Sequence: | MREVVIAEVS TQLSEVVGVI ERHLEPTLLA VHLYGSAVDG GLKPHSDIDL LVTVTVRLDE TTRRALINDL LETSASPGES EILRAVEVTI VVHDDIIPWR YPAKRELQFG EWQRNDILAG IFEPATIDID LAILLTKARE HSVALVGPAA EELFDPVPEQ DLFEALNETL TLWNSPPDWA GDERNVVLTL SRIWYSAVTG KIAPKDVAAD WAMERLPAQY QPVILEARQA YLGQEEDRLA SRADQLEEFV HYVKGEITKV VGK |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | bla OXA-1 (ARO:3001396) | Bla OXA-1 | In_Tn1935 | 291 | 12437-13312 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | OXA beta-lactamase (ARO:3000017) | ||||||||||
| Target: | penam (ARO:3000008)||carbapenem (ARO:0000020)||cephalosporin (ARO:0000032) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3001396 (bitscore: 564 )||Synonyms: | ||||||||||
| Protein Sequence: | MLAVKIKPFT KPILIMKNTI HINFAIFLII ANIIYSSASA STDISTVASP LFEGTEGCFL LYDASTNAEI AQFNKAKCAT QMAPDSTFKI ALSLMAFDAE IIDQKTIFKW DKTPKGMEIW NSNHTPKTWM QFSVVWVSQE ITQKIGLNKI KNYLKDFDYG NQDFSGDKER NNGLTEAWLE SSLKISPEEQ IQFLRKIINH NLPVKNSAIE NTIENMYLQD LDNSTKLYGK TGAGFTANRT LQNGWFEGFI ISKSGHKYVF VSALTGNLGS NLTSSIKAKK NAITILNTLN L |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI1 | IntI1 | In_Tn1935 | 337 | 13477-14490 | + |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 1 | ||||||||||
| Sequence Family: | Class 1 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MKTATAPLPP LRSVKVLDQL RERIRYLHYS LRTEQAYVHW VRAFIRFHGV RHPATLGSSE VEAFLSWLAN ERKVSVSTHR QALAALLFFY GKVLCTDLPW LQEIGRPRPS RRLPVVLTPD EVVRILGFLE GEHRLFAQLL YGTGMRISEG LQLRVKDLDF DHGTIIVREG KGSKDRALML PESLAPSLRE QLSRARAWWL KDQAEGRSGV ALPDALERKY PRAGHSWPWF WVFAQHTHST DPRSGVVRRH HMYDQTFQRA FKRAVEQAGI TKPATPHTLR HSFATALLRS GYDIRTVQDL LGHSDVSTTM IYTHVLKVGG AGVRSPLDAL PPLTSER |
||||||||||
| TnCentral Accession | TE Name | Type | Coordinates | Strand | Length |
|---|---|---|---|---|---|
| Tn4352-M20306 | Tn4352 | transposon | 1394-4073 | + | 2680 |
| IS26-MH257753 | IS26 | insertion sequence | 1394-2213 | + | 820 |
| IS26-MH257753 | IS26 | insertion sequence | 3254-4073 | + | 820 |
| IS1326_IS1353-AF071413 | IS1326::IS1353 | insertion sequence | 5367-9452 | + | 4086 |
| IS1353-AF071413 | IS1353 | insertion sequence | 5395-7008 | + | 1614 |
| Name | Associated TE | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|---|
| IR | IS26 | 1394-1407 | + | 14 | GGCACTGTTG CAAA |
| IR | IS26 | 2200-2213 | - | 14 | GGCACTGTTG CAAA |
| IR | IS26 | 3254-3267 | + | 14 | GGCACTGTTG CAAA |
| IR | IS26 | 4060-4073 | - | 14 | GGCACTGTTG CAAA |
| IRR | IS1353 | 5395-5407 | + | 13 | TGGGGGTGCG GAC |
| IRL | IS1353 | 6997-7008 | - | 12 | TGGGGTGCGG AC |