Mobile element type:
Integron
Name:
In31
Synonyms:
Accession:
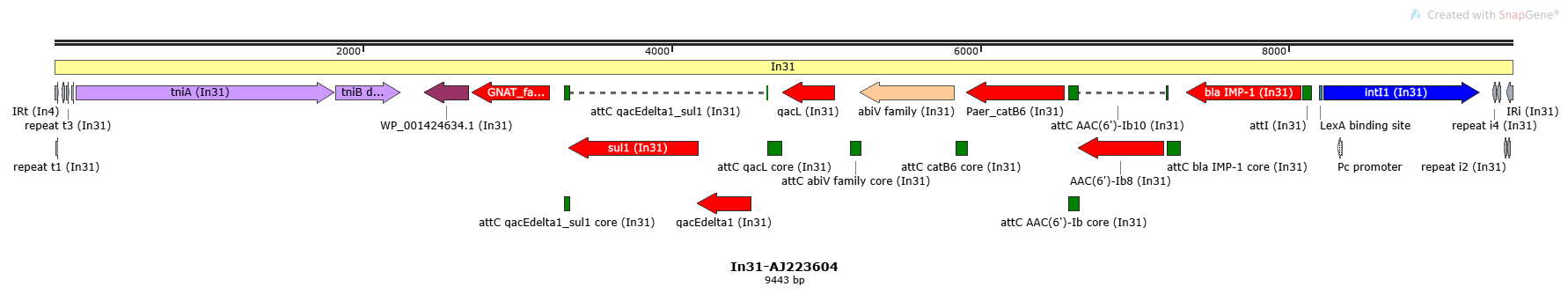
In31-AJ223604
Family:
Tn402
Group:
Class 1
First isolate:
Yes
Partial:
ND
Evidence of transposition:
ND
Host Organism:
Pseudomonas aeuroginosa 101/1477
Date of Isolation:
1999
Country:
Japan
| Name | Coordinates | Direction | Length | Sequence |
|---|---|---|---|---|
| repeat t1 | 9-27 | + | 19 | TCAGAAGACG ACTGCACCA |
| repeat t2 | 49-67 | - | 19 | TCAGGAGCTG GCTGCACAA |
| repeat t3 | 78-97 | + | 20 | TCAGAAGTGA TCTGCACCAA |
| repeat t4 | 110-128 | + | 19 | TCAATACTCG TGTGCACCA |
| repeat i4 | 9324-9342 | - | 19 | TCAGCGGACG CAGGGAGGA |
| repeat i3 | 9352-9370 | - | 19 | TCAGGCAACG ACGGGCTGC |
| repeat i2 | 9394-9412 | - | 19 | TCAGAAGCCG ACTGCACTA |
| IRi | 9411-9443 | - | 33 | TGTCGTTTTC AGAAGACGGC TGCACTGAAC GTC |
| repeat i1 | 9417-9435 | - | 19 | TCAGAAGACG GCTGCACTG |
| Name | Associated TE | Coordinates | Orientation | Length | Sequence |
|---|---|---|---|---|---|
| attC qacH_sul1 | In31 | 3307-3340 , 4618-4624 | + | 41 | CCGCTAGCGG GCGGCCGGAA GGTGAATGCT AGGCATCTAA C |
| attC qacEdelta1_sul1 core | In31 | 3307-3340 | + | 34 | CCGCTAGCGG GCGGCCGGAA GGTGAATGCT AGGC |
| attC qacL core | In31 | 4624-4711 | + | 88 | CGTTTGACGT GAGGGGCCGC CGTACCGGAA CCGAAGGGAA ACCCGCAAGC GCACCTTGGC CGGGTCCCTC TCGACGGAAT GGTTAGAT |
| attC abiV family core | In31 | 5156-5225 | + | 70 | CGTTTGGTTA AGGGGCGGCT TTAGCCCGTC CCAGCGAGCG GAGCGAGCGA TTTGAACCAT TTGTTAGGAA |
| attC catB6 core | In31 | 5845-5915 | + | 71 | CGCCCGCCAC ACGGGCGAGC AACTTGTTGC GAGTCCAGCG CCGCAGGCGA TGGTGCTGGC GCTTGTTAGA C |
| attC AAC(6')-Ib10 | In31 | 6575-6640 , 7208-7214 | + | 73 | CGTTTGACAT GAGGGGCGGC CAAGGGCGCC AGCCCTTGGA CGTCCCCCTC GATGGAAGGG TTAGGCGCCT AAC |
| attC AAC(6')-Ib core | In31 | 6575-6640 | + | 66 | CGTTTGACAT GAGGGGCGGC CAAGGGCGCC AGCCCTTGGA CGTCCCCCTC GATGGAAGGG TTAGGC |
| attC bla IMP-1 core | In31 | 7214-7298 | + | 85 | CGCTAAGTTC AGCGGCAGTT TGTAAGTTGC GCGTTGTGGA ATACTTTGCG AAGCAAACCA CAAAAGCGCA ACTTACAAAC TGTCC |
| attI | In31 | 8092-8147 | + | 56 | CTTTGTTTTA GGGCGACTGC CCTGCTGCGT AACATCGTTG CTGCTCCATA ACATCA |
ORF Summary
| Name | Associated TE | Coordinates | Orientation | Class | Subclass |
|---|---|---|---|---|---|
| tniA | In31 | 142-1821 | + | Transposase | |
| tniB delta2 | In31 | 1824-2249 | + | Accessory Gene | |
| WP_001424634.1 | In31 | 2396-2683 | - | Passenger Gene | Hypothetical |
| GNAT_fam | In31 | 2707-3207 | - | Passenger Gene | Antibiotic Resistance |
| sul1 (ARO:3000410) | In31 | 3335-4174 | - | Passenger Gene | Antibiotic Resistance |
| qacEdelta1 (ARO:3005010) | In31 | 4168-4515 | - | Passenger Gene | Antibiotic Resistance |
| qacL (ARO:3005098) | In31 | 4721-5053 | - | Passenger Gene | Antibiotic Resistance |
| abiV family | In31 | 5219-5833 | - | Passenger Gene | Other |
| Paer_catB6 (ARO:3002678) | In31 | 5910-6542 | - | Passenger Gene | Antibiotic Resistance |
| AAC(6')-Ib8 (ARO:3002579) | In31 | 6635-7186 | - | Passenger Gene | Antibiotic Resistance |
| bla IMP-1 (ARO:3002192) | In31 | 7335-8075 | - | Passenger Gene | Antibiotic Resistance |
| intI1 | In31 | 8228-9241 | + | Integron Integrase | Class 1 |
ORF Details
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniA | TniA | In31 | 559 | 142-1821 | + |
|---|
| Class: | Transposase | ||||||||||
| Transposase Chemistry: | DDE | ||||||||||
| Comment: | can be extended upstream by 12 amino acids| identical to tniA (Tn1721 and In2)| 25% amino acid sequence identity to TnsB from Tn7 | ||||||||||
| Protein Sequence: | MATDTPRIPE QGVATLPDEA WERARRRAEI ISPLAQSETV GHDAADMAAQ ALGLSRRQVY VLIRRARQGS GLVTDLVPGQ SGGGKGKGRL PEPVERVIHE LLQKRFLTKQ KRSLAAFHRE VTQVCKAQKL RVPARNTVAL RIASLDPRKV IRRREGQDAA RDLQGVGGEP PAVTAPLEQV QIDHTVIDLI VVDDRDRQPI GRPYLTLAID VFTRCVLGMV VTLEAPSAVS VGLCLVHVAC DKRPWLEGLN VEMDWQMSGK PLLLYLDNAA EFKSEALRRG CEQHGIRLDY RPLGQPHYGG IVERIIGTAM QMIHDELPGT TFSNPDQRGD YDSENKAALT LRELERWLTL AVGTYHGSVH NGLLQPPAAR WAEAVARVGV PAVVTRATSF LVDFLPILRR TLTRTGFVID HIHYYADALK PWIARRERWP SFLIRRDPRD ISRIWVLEPE GQHYLEIPYR TLSHPAVTLW EQRQALAKLR QQGREQVDES ALFRMIGQMR EIVTSAQKAT RKARRDADRR QHLKTSARPD KPVPPDTDIA DPQADNLPPA KPFDQIEEW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | tniB delta2 | TniB delta2 | In31 | 141 | 1824-2249 | + |
|---|
| Class: | Accessory Gene | ||||||||||
| Sequence Family: | ATP binding protein | ||||||||||
| Comment: | similar function to Tn7 tnsC and MuB | ||||||||||
| Protein Sequence: | MDEYPIIDLS HLLPAAQGLA RLPADERIQR LRADRWIGYP RAVEALNRLE ALYAWPNKQR MPNLLLVGPT NNGKSMIVEK FRRTHPASSD ADQEHIPVLV VQMPSEPSVI RFYVALLAAM GAPLRPRPRL PEMEQLALAW V |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | WP_001424634.1 | WP_001424634.1 | In31 | 95 | 2396-2683 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Hypothetical | ||||||||||
| Protein Sequence: | MSGWDGPVAS RSLAVRPRCL GCARGCRYPP LPSSTLDNAA AVIAADARAG LFGPVRLARP SACLGEISGG RINGGASPPF VRVKAEDSSL KNIRL |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | GNAT_fam | GNAT_fam | In31 | 166 | 2707-3207 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Sequence Family: | Acetyltransf_1 (Pfam:PF00583) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | putative acetyltransferase ADU64769.1 | ||||||||||
| Protein Sequence: | MDSEEPPNVR VACSGDIDEV VRLMHDAAAW MSAKGTPAWD VARIDRTFAE TFVLRSELLV ASCSDGIVGC CTLSAEDPEF WPDALKGEAA YLHKLAVRRT HAGRGVSSAL IEACRHAART QGCAKLRLDC HPNLRGLYER LGFTHVDTFN PGWDPTFIAE RLELEI |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | sul1 (ARO:3000410) | Sul1 | In31 | 279 | 3335-4174 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic target replacement (ARO:0001002) | ||||||||||
| Sequence Family: | sulfonamide resistant sul (ARO:3004238) | ||||||||||
| Transposase Chemistry: | dihydropteroate synthase | ||||||||||
| Target: | sulfonamide antibiotic (ARO:3000282)||sulfone antibiotic (ARO:3003401) | ||||||||||
| Comment: | perfect match to reference sequence for ARO:3000410 | ||||||||||
| Protein Sequence: | MVTVFGILNL TEDSFFDESR RLDPAGAVTA AIEMLRVGSD VVDVGPAASH PDARPVSPAD EIRRIAPLLD ALSDQMHRVS IDSFQPETQR YALKRGVGYL NDIQGFPDPA LYPDIAEADC RLVVMHSAQR DGIATRTGHL RPEDALDEIV RFFEARVSAL RRSGVAADRL ILDPGMGFFL SPAPETSLHV LSNLQKLKSA LGLPLLVSVS RKSFLGATVG LPVKDLGPAS LAAELHAIGN GADYVRTHAP GDLRSAITFS ETLAKFRSRD ARDRGLDHA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacEdelta1 (ARO:3005010) | QacEdelta1 | In31 | 115 | 4168-4515 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | major facilitator superfamily (MFS) antibiotic efflux pump (ARO:0010002) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | subunit of the qac multidrug efflux pump||perfect match to reference sequence for ARO:3005010 (bitscore:219) | ||||||||||
| Protein Sequence: | MKGWLFLVIA IVGEVIATSA LKSSEGFTKL APSAVVIIGY GIAFYFLSLV LKSIPVGVAY AVWSGLGVVI ITAIAWLLHG QKLDAWGFVG MGLIIAAFLL ARSPSWKSLR RPTPW |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | qacL (ARO:3005098) | QacL | In31 | 110 | 4721-5053 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic efflux (ARO:0010000) | ||||||||||
| Sequence Family: | small multidrug resistance (SMR) antibiotic efflux pump (ARO:0010003) | ||||||||||
| Target: | disinfecting agents and antiseptics(ARO:3005386) | ||||||||||
| Comment: | subunit of the qac multidrug efflux pump||loose match to reference sequence for ARO:3005098 (bitscore:173) | ||||||||||
| Protein Sequence: | MKNWLFLATA IIFEVIATSA LKSSEGFTRL VPSFIVVAGY AAAFYFLSLT LKSIPVGIAY AVWSGLGIVL VTAIAWVLHG QKLDMWGFVG VGFIISGVAV LNLLSKASVH |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | abiV family | AbiV family | In31 | 204 | 5219-5833 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Other | ||||||||||
| Sequence Family: | AbiV family abortive infection protein (WP_017363802.1) | ||||||||||
| Protein Sequence: | MDVDLQSIGI QTVQEALEKG SLIFNEKTKT DDFNRACDHI LVLLVDAFQC FDRGSWGTSV FLSITAIEEV AKAEVGLYRR EGKIGKAKRG KDKLFNHQEK HRMAILPTVF MSKRLEEALG KEKCAELLKE AAHGEFRNLR ESSLYFSNEN GQFVTPANVV SQNRAKEFLL LALEAADDRL VGYTNHTGIL EAKINEIFSV VAHS |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | Paer_catB6 (ARO:3002678) | Paer_catB6 | In31 | 210 | 5910-6542 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | chloramphenicol acetyltransferase (CAT) (ARO:3000122) | ||||||||||
| Target: | phenicol antibiotic (ARO:3000387) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002678 (bitscore: 438) | ||||||||||
| Protein Sequence: | MENYFDSPFK GKLLSEQVTN RNIKVGRYSY YSGYYHGHSF DDCARYLLPD RDDVDKLIIG SFCSIGSGAS FIMAGNQGHR HDWVTSFPFF YMQEEPAFSS STDAFQKAGD TIVGNDVWIG SEAMIMPGIK IGDGAVIGSR SLVTRDVEPY TIIGGNPAKQ IKKRFSDEEI SLLMEMEWWN WPLDKIKTAM PLLCSSDIFG LHRHWRGIAV |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | AAC(6')-Ib8 (ARO:3002579) | AAC(6')-Ib8 | In31 | 183 | 6635-7186 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | AAC(6') (ARO:3000345) | ||||||||||
| Target: | aminoglycoside antibiotic (ARO:0000016) | ||||||||||
| Comment: | strict match to reference sequence for ARO:3002579 missing 43 aa N-ter (bitscore: 379) | ||||||||||
| Protein Sequence: | TNSTDSVTLR LMTEHDLAML YEWLNRSHIV EWWGGEEARP TLADVQEQYL PSVLAQESVT PYIAMLNGEP IGYAQSYVAL GSGDGWWEEE TDPGVRGIDQ LLANASQLGK GLGTKLVRAL VELLFNDPEV TKIQTDPSPS NLRAIRCYEK AGFERQGTVT TPDGPAVYMV QTRQAFERTR SDA |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | bla IMP-1 (ARO:3002192) | Bla IMP-1 | In31 | 246 | 7335-8075 | - |
|---|
| Class: | Passenger Gene | ||||||||||
| Subclass: | Antibiotic Resistance | ||||||||||
| Function: | antibiotic inactivation (ARO:0001004) | ||||||||||
| Sequence Family: | IMP beta-lactamase (ARO:3000020) | ||||||||||
| Target: | cephalosporin (ARO:0000032)||penem (ARO:3003706)||penam (ARO:3000008)||cephamycin (ARO:0000044)||carbapenem (ARO:0000020) | ||||||||||
| Comment: | 100% identical to reference sequence forARO:3002192 | ||||||||||
| Protein Sequence: | MSKLSVFFIF LFCSIATAAE SLPDLKIEKL DEGVYVHTSF EEVNGWGVVP KHGLVVLVNA EAYLIDTPFT AKDTEKLVTW FVERGYKIKG SISSHFHSDS TGGIEWLNSR SIPTYASELT NELLKKDGKV QATNSFSGVN YWLVKNKIEV FYPGPGHTPD NVVVWLPERK ILFGGCFIKP YGLGNLGDAN IEAWPKSAKL LKSKYGKAKL VVPSHSEVGD ASLLKLTLEQ AVKGLNESKK PSKPSN |
||||||||||
| Gene Name | Protein Name | Associated TE | Gene Length | Coordinates | Strand | intI1 | IntI1 | In31 | 337 | 8228-9241 | + |
|---|
| Class: | Integron Integrase | ||||||||||
| Subclass: | Class 1 | ||||||||||
| Sequence Family: | Class 1 Integron Tyrosine Integrase | ||||||||||
| Transposase Chemistry: | Tyrosine | ||||||||||
| Protein Sequence: | MKTATAPLPP LRSVKVLDQL RERIRYLHYS LRTEQAYVHW VRAFIRFHGV RHPATLGSSE VEAFLSWLAN ERKVSVSTHR QALAALLFFY GKVLCTDLPW LQEIGRPRPS RRLPVVLTPD EVVRILGFLE GEHRLFAQLL YGTGMRISEG LQLRVKDLDF DHGTIIVREG KGSKDRALML PESLAPTLRE QLSRARAWWL KDQAEGRSGV ALPDALERKY PRAGHSWPWF WVFAQHTHST DPRSGVVRRH HMYDQTFQRA FKRAVEQAGI TKPATPHTLR HSFATALLRS GYDIRTVQDL LGHSDVSTTM IYTHVLKVGG AGVRSPLDAL PPLTSER |
||||||||||
1. Laraki N, Galleni M, Thamm I, Riccio ML, Amicosante G, Frere JM, Rossolini GM.
Structure of In31, a blaIMP-containing Pseudomonas aeruginosa integron phyletically related to In5, which carries an unusual array of gene cassettes.. Antimicrob Agents Chemother. 1999 Apr;43(4):890-901. doi: 10.1128/AAC.43.4.890. PubMed ID:
10103196
.