Difference between revisions of "IS Families/ISAzo13 family"
Jump to navigation
Jump to search
| Line 5: | Line 5: | ||
<br /> | <br /> | ||
<hr> | <hr> | ||
| + | {{TnPedia}} | ||
Latest revision as of 12:55, 5 December 2022
This family, represented by 37 members in ISfinder emerged from the ISNCY orphan group. It is based on both Tpase and IR sequence similarities (Fig.ISAzo13). Insertion generates a 3bp AT-rich DR and the ends have a consensus GGa/g. Their Tpases are highly conserved with a probable DDE motif and an HTH motif at the N-terminus which could function as a DNA binding domain. Two members encode two orfs with a possible PRF (Programmed Transcriptional Frameshifting) motif of 8 or 9 A while the other members encode a unique orf which includes a triple lysine at the equivalent position (Gourbeyre, unpublished)[1].

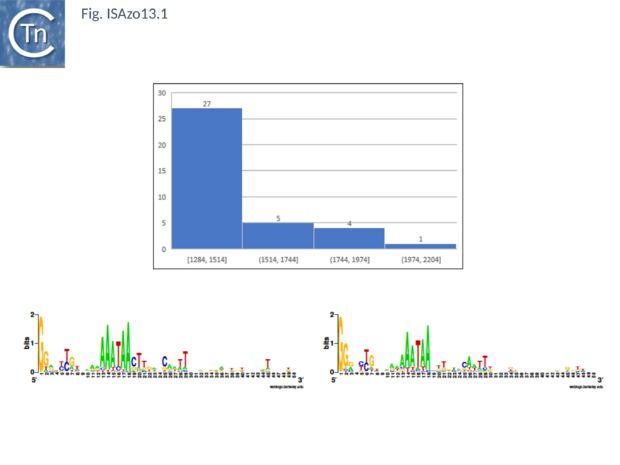
Fig. ISAzo13. General ISAzo13 characteristics, average length and common ends. Top: Distribution of IS length (base pairs) of ISAzo13 family members. The number of examples used in the sample is shown above each column. Bottom: Left (IRL) and right IRR inverted terminal repeats are shown in WebLogo format.
Bibliography
- ↑ Siguier P, Perochon J, Lestrade L, Mahillon J, Chandler M . ISfinder: the reference centre for bacterial insertion sequences. - Nucleic Acids Res: 2006 Jan 1, 34(Database issue);D32-6 [PubMed:16381877] [DOI]