Difference between revisions of "General Information/ IS derivatives of Tn3 family transposons"
(Created page with "Another source of ambiguity for classification purposes occurs in the Tn3 family (see section “Tn families”)<ref><nowiki><pubmed>10477306</pubmed></nowiki></ref>(Fig.1.15.1 and 1.15.2). Tn...") |
|||
| Line 1: | Line 1: | ||
| − | Another source of ambiguity for classification purposes occurs in the Tn3 family (see section “Tn families”)<ref><nowiki><pubmed>10477306</pubmed></nowiki></ref>(Fig.1.15.1 and 1.15.2). Tn3 family members are quite variable. They include a number of diverse passenger genes | + | Another source of ambiguity for classification purposes occurs in the Tn3 family (see section “Tn families”)<ref><nowiki><pubmed>10477306</pubmed></nowiki></ref> [[:Image:1.15.1.png|(Fig.1.15.1]] and [[:Image:1.15.1.png| 1.15.2)]]. Tn3 family members are quite variable. They include a number of diverse passenger genes that can represent entire operons, notably mercury resistance, or individual genes involved in antibiotic resistance, breakdown of halogenated aromatics, or virulence [e.g. <ref><nowiki><pubmed>10477306</pubmed></nowiki></ref>]]. They often carry integron recombination platforms enabling them to incorporate additional resistance genes by recruiting integron cassettes<ref><nowiki><pubmed>16845431</pubmed></nowiki></ref>. Members are quite characteristic: they have long relatively well conserved IR and a particularly long Tpase (950 to 1025 aa). They also encode a site-specific recombination (“resolution”) system necessary for completion of their transposition<ref><nowiki><pubmed>10477306</pubmed></nowiki></ref>. There are a number of different resolution systems associated with different members of this family [[:Image:1.15.2.png|(Fig.1.15.2)]] IS1071, [[:Image:1.15.1.png|(Fig.1.15.1)]] composed of Tn3-like IR and Tpase gene but lacking both the site-specific recombination system and passenger genes was identified many years ago<ref><nowiki><pubmed>1656436</pubmed></nowiki></ref><ref><nowiki><pubmed>8117095</pubmed></nowiki></ref>. This clearly accords with the definition of an IS. Several other examples have now been identified (e.g. ISVsa19, ISShfr9, ISBusp1). |
[[Image:1.15.1.png|thumb|center|600px|Fig 1.15.1]] | [[Image:1.15.1.png|thumb|center|600px|Fig 1.15.1]] | ||
[[Image:1.15.2.png|thumb|center|600px|Fig 1.15.2]] | [[Image:1.15.2.png|thumb|center|600px|Fig 1.15.2]] | ||
| − | + | ==Bibliography== | |
<references/> | <references/> | ||
| − | |||
| − | |||
Revision as of 17:52, 30 April 2020
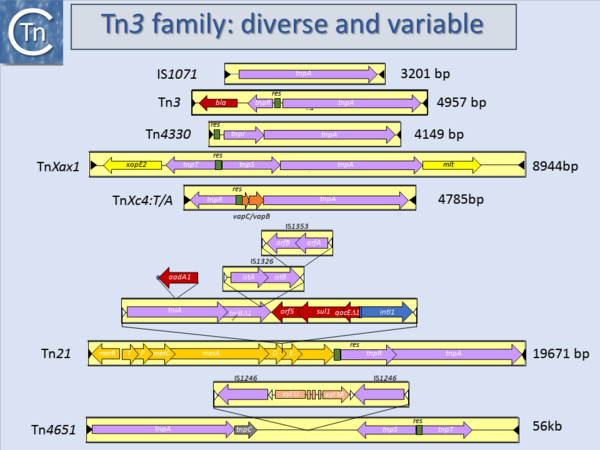
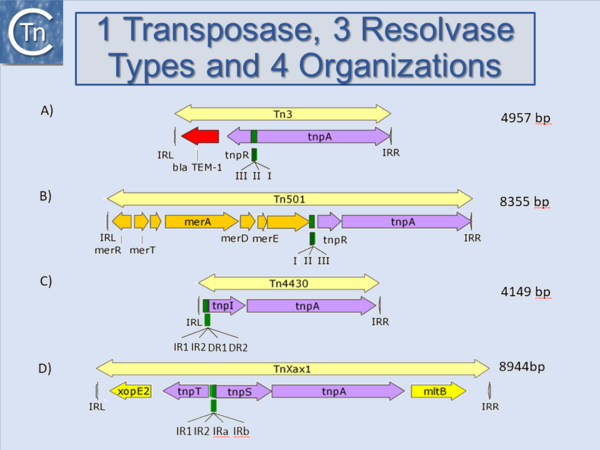
Another source of ambiguity for classification purposes occurs in the Tn3 family (see section “Tn families”)[1] (Fig.1.15.1 and 1.15.2). Tn3 family members are quite variable. They include a number of diverse passenger genes that can represent entire operons, notably mercury resistance, or individual genes involved in antibiotic resistance, breakdown of halogenated aromatics, or virulence [e.g. [2]]]. They often carry integron recombination platforms enabling them to incorporate additional resistance genes by recruiting integron cassettes[3]. Members are quite characteristic: they have long relatively well conserved IR and a particularly long Tpase (950 to 1025 aa). They also encode a site-specific recombination (“resolution”) system necessary for completion of their transposition[4]. There are a number of different resolution systems associated with different members of this family (Fig.1.15.2) IS1071, (Fig.1.15.1) composed of Tn3-like IR and Tpase gene but lacking both the site-specific recombination system and passenger genes was identified many years ago[5][6]. This clearly accords with the definition of an IS. Several other examples have now been identified (e.g. ISVsa19, ISShfr9, ISBusp1).