Difference between revisions of "General Information/Major Groups are Defined by the Type of Transposase They Use"
| Line 187: | Line 187: | ||
| + | |||
| + | |||
| + | |||
| + | <br /> | ||
==Groups with S-Transposases== | ==Groups with S-Transposases== | ||
Revision as of 12:16, 4 May 2020
The principal factor in IS classification is the similarity, at the primary sequence level, of the enzymes which catalyze their movement, their transposases (Tpases). In addition, a variety of characteristics are also taken into account. These include: the length and sequence of the short imperfect terminal inverted repeat sequences (IRs) carried by many ISs at their ends (TIRs or ITRs in eukaryotes); the length and sequence of the short flanking direct target DNA repeats (DRs) (TSD, Target Site Duplication, in eukaryotes) often generated on insertion; the organization of their open reading frames or the target sequences into which they insert[1][2][3]. IS and some transposons can also be divided into two major types based on the chemistry used in breaking and rejoining DNA during TE displacement: the DDE (and DEDD) and HUH enzymes. Additional types of transposase enzymes have been identified (Fig.1.7.1) but are generally associated with other types of transposon rather than IS.
A relatively new type of potential transposase, Cas1, is associated with so-called casposons, elements that may resemble complex IS and are related to CRISPRs (for more details please see The Casposases section).
Contents
Groups with DDE Transposases
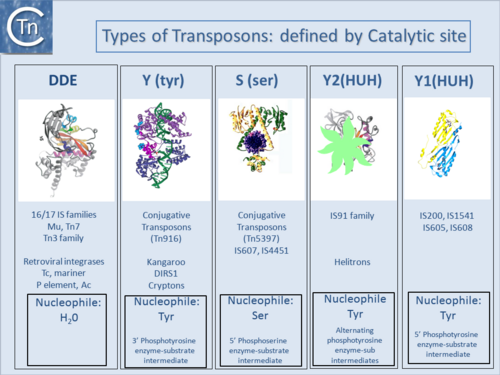
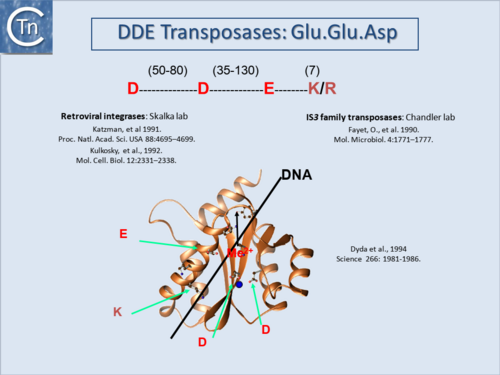
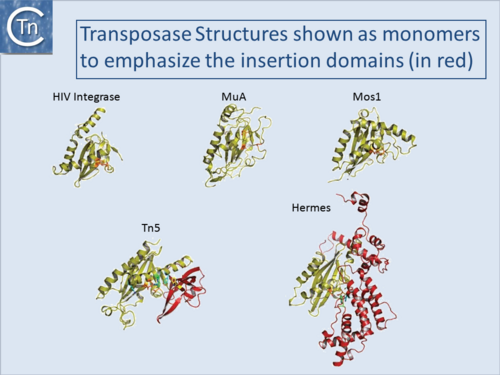
DDE enzymes, so-called because of a conserved Asp, Asp, Glu triad of amino acids which coordinate essential metal ions, use OH (e.g. H20) as a nucleophile in a transesterification reaction[4] (Fig.1.7.1) and (Fig.1.8.1). IS with DDE enzymes are the most abundant type in the public databases (Fig.1.4.2). This is partly due to the fact that the definition of an IS became implicitly coupled to the presence of a DDE Tpase, an idea probably reinforced by the similarity between Tpases of IS (and other TE) and the retroviral integrases (Fig.1.8.1)[5][6][7] particularly in the region including the catalytic site. More precisely, for these TE, the triad is DD(35)E in which the second D and E are separated by 35 residues. As more DDE transposases were identified, the distance separating the D and E residues was found to vary slightly (TABLE MGE transposases examined using secondary structure prediction programmes)[8]. However, for certain IS, this distance was significantly larger. In these cases, the Tpases include an “insertion domain” between the second D and E residues[9] with either α-helical or β-strand configurations (Fig.1.8.2). Although in most cases this is a prediction, it has been confirmed by crystallographic studies for the IS50 [β-strand[10] and Hermes [α-helical;[11] Tpases. The function of these “insertion domains” is not entirely clear[12].
Transposases examined by secondary structure prediction programs
| Table 2. Adapted from Hickman et al. 2010, Integrating prokaryotes and eukaryotes: DNA transposases in light of structure. 1Information on the number of copies within the host genome was obtained from ISfinder or the reference indicated by the asterisk. 2Where indicated, the secondary structure predicts an insertion domain between β5 and α4 with predominantly either β-strands or α-helices. 3Relevant references include reviews or papers that report the results of secondary structure prediction, report sequence alignments or consensus sequences, identify the DDE/D catalytic residues, or demonstrate that the element is active. The association of certain eukaryotic superfamilies to specific IS families is as per Feschotte and Pritham (2005) and references therein. | ||||||||
| Family | Element (or protein) analyzed | Active or # copies in genome1 | From secondary structure, type of DDE/D motif2 | Relevant references3 | ||||
|---|---|---|---|---|---|---|---|---|
| IS1 | IS1N | >40* | DD(24)E | *Nyman et al., 1981; Ohta et al., 2002, 2004; Siguier et al., 2009 | ||||
| ISSto9 | 5 | DD(20)E | ||||||
| IS1595 | ISPna2 | — | DD(36)N | Siguier et al., 2009 | ||||
| ISH4 | — | DD(36)E | ||||||
| IS1016C | — | DD(34)E | ||||||
| IS1595 | — | DD(35)N | ||||||
| ISSod11 | 13 | DD(34)H | ||||||
| ISNWi1 | — | DD(35)E | ||||||
| ISNha5 | — | DD(33)E | ||||||
| Merlin: MERLIN1_SM | consensus | DD(36)E | Feschotte, 2004 | |||||
| IS3 | IS911 | Active | DD(35)E | Polard and Chandler, 1995; Rousseau et al., 2002 | ||||
| IS481 | IS481 | ~100* | DD(35)E | *Glare et al., 1990; Chandler and Mahillon, 2002 | ||||
| IS4 | IS50R | Active | PDB ID: 1muh | Rezsöhazy et al., 1993; Davies et al., 2000 | ||||
| IS701 | IS701 | Active (15*) | DD(β-strand)E | *Mazel et al., 1991 | ||||
| ISRso17 | 7 | |||||||
| ISH3 | ISC1359 | 5 | DD(β-strand)E | — | ||||
| ISC1439A | 13 | |||||||
| IS1634 | IS1634 | Active (~30*) | DD(β-strand)E | *Vilei et al., 1999 | ||||
| ISMac5 | 7 | |||||||
| ISPlu4 | 7 | |||||||
| IS5 | IS903 | Active | DD(65)E | Derbyshire et al., 1987; Rezsöhazy et al., 1993; Tavakoli et al., 1997 | ||||
| PIF/Harbinger: PIFa (Z. mays) | Active | DD(59)E | Zhang et al., 2001; Kapitonov and Jurka, 2004; Sinzelle et al., 2008 | |||||
| IS1182 | "IS660 | 3 | DD(β-strand)E | Takami et al., 2001 | ||||
| ISPsy6 | 14 | |||||||
| IS6 | IS6100 | Active | DD(34)E | Martin et al., 1990; Mahillon and Chandler, 1998 | ||||
| IS21 | IS21 | Active | DD(45)E | Mahillon and Chandler, 1998; Berger and Haas, 2001 | ||||
| IS30 | IS30 | Active | DD(33)E | Caspers et al., 1984; Mahillon and Chandler, 1998 | ||||
| IS66 | IS679 | Active | DD(α-helical?)E | Han et al., 2001 | ||||
| ISPsy5 | 33 | |||||||
| ISMac8 | 3 | |||||||
| IS110 | IS492 | Active | DEDD | Perkins-Balding et al., 1999; Buchner et al., 2005 | ||||
| IS1111 | 20 | |||||||
| IS256 | IS256 | Active | DD(α-helical)E | Mahillon and Chandler, 1998; Prudhomme et al., 2002 | ||||
| MuDr/Foldback</i< (Mutator) | Active | Eisen et al., 1994; Babu et al., 2006; Hua-Van and Capy, 2008 | ||||||
| IS630 | ISY100 | Active | DD(34)E | Doak et al., 1994; Feng and Colloms, 2007 | ||||
| Tc1/mariner: Mos1 (D. mauritiana) | PDB ID: 2f7t | Plasterk et al., 1999; Richardson et al., 2006 | ||||||
| Zator: Zator-1_HM | 36* | DD(43)E | *Bao et al., 2009 | |||||
| IS982 | ISPfu3 | 5 | DD(47)E | Mahillon and Chandler, 1998 | ||||
| IS1380 | IS1380A | ~100* | DD(β-strand)E | *Takemura et al., 1991; Chandler and Mahillon, 2002 | ||||
| piggyBac (T. ni) | Active | DD(β-strand)D | Cary et al., 1989; Sarkar et al., 2003; Mitra et al., 2008 | |||||
| ISAs1 | ISAzo3 | 7 | DD(β-strand)E/D? | — | ||||
| ISL3 | IS31831 | Active | DD(α-helical)E | Suzuki et al., 2006 | ||||
| IS651 | 22 | |||||||
| Tn3 | Tn3 (E. coli) | Active | DD(α-helical?)E, DD(α-helical)E insertion | Grindley, 2002 | ||||
| hAT | Hermes | Active | PDB ID: 2bw3 | Warren et al., 1994; Rubin et al., 2001; Hickman et al., 2005 | ||||
| CACTA | CACTA1 (A. thaliana) En/Spm ZM | Active | DD(α-helical?)E/D? | Miura et al., 2001; DeMarco et al., 2006 | ||||
| P | Drosophila | Active | ? | Rio, 2002 | ||||
| Transib | >Transib1_AG | Consensus | DD(α-helical)E | Kapitonov and Jurka, 2005; Chen and Li, 2008 | ||||
| RAG1 (M. musculus) | Active | Kim et al., 1999; Landree et al., 1999; Lu et al., 2006 | ||||||
| Sola | Sola3-3_HM | Multiple copies* | DD(40)E | *Bao et al., 2009 | ||||
Major DDE transposition pathways
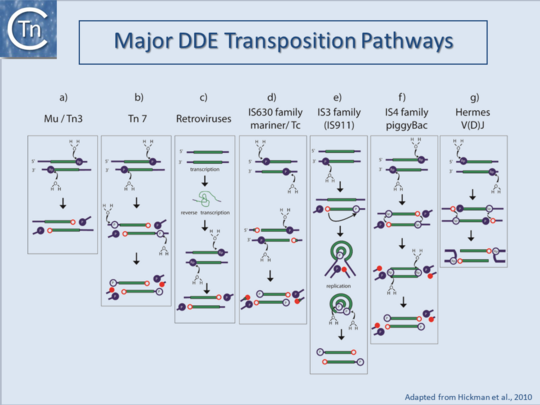
Although DDE-type transposons share basic transposition chemistry, different TE vary in the steps leading to the formation of a unique insertion intermediate (Fig.1.8.3)[13][14]. They catalyze the cleavage of a single DNA strand to generate a 3’OH at the TE ends which is subsequently used as a nucleophile to attack the DNA target phosphate backbone. This is known as the transferred strand. The variations are due to the way in which the second (non-transferred) strand is processed[15][16][17].
There are several ways in which second-strand processing can occur (Fig.1.8.3): for certain IS, the second strand is not cleaved but replication following the transfer of the first strand fuses donor and target molecules to generate cointegrates with a directly repeated copy at each donor/target junction. This is known as replicative transposition (e.g. IS6, Tn3) or more precisely, Target Primed Replicative Transposition (TPRT) (Fig.1.8.3 pathway a).
In the other pathways, the flanking donor DNA can be shed in several different ways: the non-transferred strand may be cleaved initially several bases within the IS prior to cleavage of the transferred strand [e.g. IS630 and Tc1[18][19][20] (Fig.1.8.3 pathway d); the 3’OH generated by the first-strand cleavage may be used to attack the second strand to form a hairpin structure at the IS ends liberating the IS from flanking DNA and subsequently hydrolyzed to regenerate the 3’OH known as conservative or cut-and-paste transposition (e.g. IS4;[21] (Fig.1.8.3 pathway f) and (Tn10 movie - see below) (IS4.4; IS4.5; IS4.6; IS4.7); the 3’OH of the transferred strand from one IS end may attack the other to generate a donor molecule with a single strand bridge which is then replicated to produce a double-strand transposon circle intermediate and regenerating the original donor molecule known as copy-out-paste-in or more precisely Donor Primed Replicative Transposition (DPRT) (e.g. IS3 ) [22] (Fig.1.8.2 pathway e) and (IS911 movie - see below); or the 3’OH at the flank of the non-transferred strand may attack the second strand to form a hairpin on the flanking DNA and a 3’OH on the transferred strand (at present this has only been demonstrated for eukaryotic TE of the hAT family and in V(D)J recombination [23]) (Fig.1.8.3 pathway g).
Clearly, many families produce double-strand circular intermediates but this does not necessarily mean that they all use the copy-paste DPRT mechanism since a circle could formally be generated by excision involving recombination of both strands[24]. These differences are reflected in the different IS families.
IS911.Copy out - Paste in (column e in figure 1.8.3)
|
Tn10. Cut and paste (column f in figure 1.8.3)
|
|---|
Groups with DEDD Transposases
A similar type of Tpase, known as a DEDD Tpase, is related to the Holiday junction resolvase, RuvC (Choi, et al., 2003, Buchner, et al., 2005)[25][26][27] but is at present limited to only a single known IS family (IS110). The organization of family members is quite different from that of the DDE ISs: they do not contain the typical terminal IRs of the DDE IS (although one subgroup, IS1111, carry sub-terminal IR) and do not generate flanking target DRs on insertion. This implies that their transposition occurs using a different mechanism to the DDE IS. It seems probable that an intermediate resembling a four-way Holliday junction is involved. Moreover, in contrast to the DDE transposases in which a DNA binding domain invariably precedes the catalytic domain, DEDD transposases appear to include a DNA binding domain downstream from the catalytic domain.
Groups with HUH Enzymes
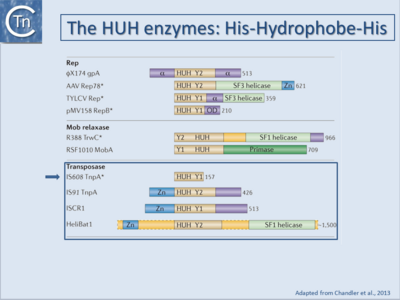
TE encoding the second major type of Tpase, called HUH (named for the conserved active site amino acid residues H=Histidine and U=large hydrophobic residue )(Fig.1.7.1) and (Fig.1.10.1), has been identified more recently. HUH enzymes are widespread single-strand nucleases. They include Rep proteins involved in bacteriophage and plasmid rolling circle replication and relaxases or Mob proteins involved in conjugative plasmid transfer[28]. They are limited to two prokaryotic (IS91 and IS200/IS605; [29]) and one eukaryotic (helitron[30]) TE family.
As Tpases, they are involved in presumed rolling circle transposition and also in single-strand transposition (see [31][32]). Not only is the transposition chemistry radically different to that of DDE group elements, since it involves DNA cleavage using a tyrosine residue and transient formation of a phospho-tyrosine bond, but the associated transposons have an entirely different organization and include sub-terminal secondary structures instead of IRs (see IS families below [33]). Note that these Tpases are not related to the well-characterized tyrosine site-specific recombinases such as phage integrases.
There are two major HUH Tpase families: Y1 and Y2 enzymes (Fig.1.7.1)(see [34]) depending on whether there is a single or two catalytic Y residues. One family includes IS91-family transposases[35][36], the other includes IS200/IS605 transposases[37][38][39][40][41][42]. Although these enzymes use the same Y-mediated cleavage mechanism, IS200/IS605 family Y1 transposases and IS91 transposases appear to carry out the transposition process in quite different ways. Neither carries terminal IRs nor do they generate DRs on insertion. Members of these families transpose using an entirely different mechanism to IS with DDE transposases[43][44]. The members of the IS91 insertion sequence family[45][46], are related to newly defined group, the ISCR[47] (see “IS91-related ISCRs”) and with eukaryotic helitrons (Fig.1.7.1)[48]. These IS carry sub-terminal sequences which are able to form hairpin secondary structures (Fig.1.3.2). This is particularly marked in the IS200/IS605 family elements and, at least in the case of this family, it is these structures which are recognised by the transposase[49].
Groups with S-Transposases
The third transposase family is represented by IS607 which carries a Tpase closely related to serine recombinases such as the resolvases of Tn3 family elements. Little is known about their transposition mechanism. However, it appears likely, in view of the known activities of resolvases, that IS607 transposition may involve a double-strand DNA intermediate (Grindley cited as pers. comm. in [50]) see also [51] (Fig.1.7.1).
Groups with Y-Transposases
Finally, tyrosine site-specific recombinases of the bacteriophage integrase (Int) type are often associated with conjugative transposons (Integrative Conjugative Elements or ICE)( IS related to ICE) and are considered to be Tpases. However, at present there are no known IS which use this type of enzyme (Fig.1.7.1).
Bibliography
- ↑ <pubmed>9729608</pubmed>
- ↑ <pubmed>24499397</pubmed>
- ↑ <pubmed>26104715</pubmed>
- ↑ <pubmed>26104718</pubmed>
- ↑ <pubmed>1963920</pubmed>
- ↑ <pubmed>1850126</pubmed>
- ↑ <pubmed>1314954</pubmed>
- ↑ <pubmed>20067338</pubmed>
- ↑ <pubmed>20067338</pubmed>
- ↑ <pubmed>10207011</pubmed>
- ↑ <pubmed>16041385</pubmed>
- ↑ <pubmed>20067338</pubmed>
- ↑ <pubmed>26104718</pubmed>
- ↑ <pubmed>20067338</pubmed>
- ↑ <pubmed>26104718</pubmed>
- ↑ <pubmed>10838584</pubmed>
- ↑ <pubmed>14682279</pubmed>
- ↑ <pubmed>8556864</pubmed>
- ↑ <pubmed>7954797</pubmed>
- ↑ <pubmed>17680987</pubmed>
- ↑ <pubmed>26104553</pubmed>
- ↑ <pubmed>26350305</pubmed>
- ↑ <pubmed>15616554</pubmed>
- ↑ <pubmed>26104718</pubmed>
- ↑ <pubmed>12897009</pubmed>
- ↑ <pubmed>15866929</pubmed>
- ↑ <pubmed>11169105</pubmed>
- ↑ <pubmed>23832240</pubmed>
- ↑ <pubmed>26350330</pubmed>
- ↑ <pubmed>26350323</pubmed>
- ↑ <pubmed>26350330</pubmed>
- ↑ <pubmed>26104718</pubmed>
- ↑ <pubmed>26350330</pubmed>
- ↑ <pubmed>23832240</pubmed>
- ↑ <pubmed>6282809</pubmed>
- ↑ <pubmed>19709290</pubmed>
- ↑ <pubmed>6315530</pubmed>
- ↑ <pubmed>3009825</pubmed>
- ↑ <pubmed>6313217</pubmed>
- ↑ <pubmed>11807059</pubmed>
- ↑ <pubmed>9631304</pubmed>
- ↑ <pubmed>9858724</pubmed>
- ↑ <pubmed>11136468</pubmed>
- ↑ <pubmed>16163392</pubmed>
- ↑ <pubmed>1321417</pubmed>
- ↑ <pubmed>1310503</pubmed>
- ↑ <pubmed>16760305</pubmed>
- ↑ <pubmed>17850916</pubmed>
- ↑ <pubmed>26350330</pubmed>
- ↑ <pubmed>17347521</pubmed>
- ↑ <pubmed>24195768</pubmed>